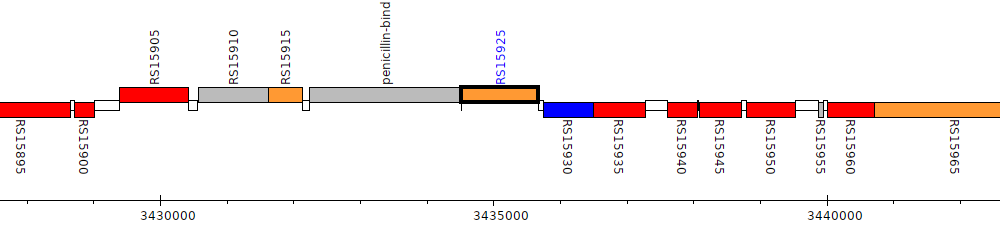
Burkholderia cenocepacia H111, I35_RS15925
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0008360 | regulation of cell shape |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02210
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0051301 | cell division |
Inferred from Sequence Model
Term mapped from: InterPro:PF01098
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02210
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Hamap | MF_02079 | Peptidoglycan glycosyltransferase MrdB [mrdB]. | IPR011923 | Probable peptidoglycan glycosyltransferase RodA/MrdB | 14 | 380 | 28.439 |
| Pfam | PF01098 | Cell cycle protein | IPR001182 | Probable peptidoglycan glycosyltransferase FtsW/RodA | 24 | 378 | 5.6E-99 |
| ProSitePatterns | PS00428 | Cell cycle proteins ftsW / rodA / spoVE signature. | IPR018365 | Cell cycle, FtsW / RodA / SpoVE, conserved site | 335 | 359 | - |
| TIGRFAM | TIGR02210 | rodA_shape: rod shape-determining protein RodA | IPR011923 | Probable peptidoglycan glycosyltransferase RodA/MrdB | 20 | 377 | 2.9E-126 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.