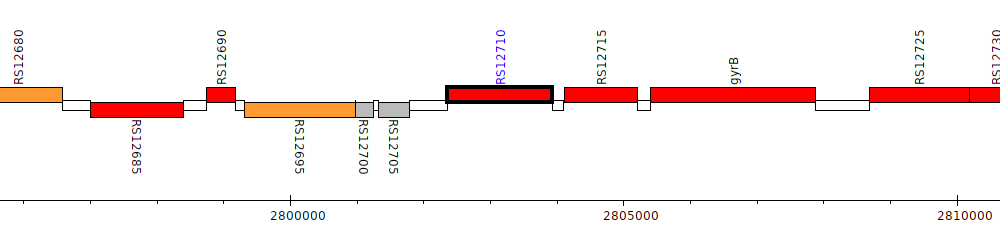
Burkholderia cenocepacia AU 1054, BCEN_RS12710
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00362
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006270 | DNA replication initiation |
Inferred from Sequence Model
Term mapped from: InterPro:cd06571
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003688 | DNA replication origin binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00362
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006275 | regulation of DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:cd06571
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd06571
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0043565 | sequence-specific DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd06571
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006270 | DNA replication initiation |
Inferred from Sequence Model
Term mapped from: InterPro:cd06571
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003688 | DNA replication origin binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00362
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0043565 | sequence-specific DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd06571
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006275 | regulation of DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:cd06571
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd06571
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00362
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Coils | Coil | 509 | 525 | - | |||
| Gene3D | G3DSA:3.30.300.180 | IPR038454 | DnaA, N-terminal domain superfamily | 2 | 103 | 1.1E-22 | |
| SMART | SM00382 | IPR003593 | AAA+ ATPase domain | 222 | 356 | 7.8E-9 | |
| PRINTS | PR00051 | Bacterial chromosomal replication initiator (DNAA) signature | IPR020591 | Chromosomal replication control, initiator DnaA-like | 223 | 243 | 1.2E-49 |
| TIGRFAM | TIGR00362 | DnaA: chromosomal replication initiator protein DnaA | IPR001957 | Chromosomal replication control, initiator DnaA | 5 | 523 | 2.0E-169 |
| Pfam | PF08299 | Bacterial dnaA protein helix-turn-helix | IPR013159 | Chromosomal replication initiator, DnaA C-terminal | 434 | 502 | 1.5E-32 |
| PRINTS | PR00051 | Bacterial chromosomal replication initiator (DNAA) signature | IPR020591 | Chromosomal replication control, initiator DnaA-like | 483 | 502 | 1.2E-49 |
| SUPERFAMILY | SSF48295 | IPR010921 | Trp repressor/replication initiator | 421 | 524 | 3.14E-38 | |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 189 | 401 | 2.53E-43 | |
| PRINTS | PR00051 | Bacterial chromosomal replication initiator (DNAA) signature | IPR020591 | Chromosomal replication control, initiator DnaA-like | 321 | 348 | 1.2E-49 |
| SMART | SM00760 | IPR013159 | Chromosomal replication initiator, DnaA C-terminal | 433 | 502 | 3.9E-44 | |
| Gene3D | G3DSA:3.40.50.300 | 188 | 355 | 6.5E-47 | |||
| PRINTS | PR00051 | Bacterial chromosomal replication initiator (DNAA) signature | IPR020591 | Chromosomal replication control, initiator DnaA-like | 255 | 269 | 1.2E-49 |
| PRINTS | PR00051 | Bacterial chromosomal replication initiator (DNAA) signature | IPR020591 | Chromosomal replication control, initiator DnaA-like | 287 | 301 | 1.2E-49 |
| Hamap | MF_00377 | Chromosomal replication initiator protein DnaA [dnaA]. | IPR001957 | Chromosomal replication control, initiator DnaA | 2 | 524 | 31.984 |
| Pfam | PF11638 | DnaA N-terminal domain | IPR024633 | DnaA N-terminal domain | 3 | 66 | 1.0E-17 |
| ProSitePatterns | PS01008 | DnaA protein signature. | IPR018312 | Chromosomal replication control, initiator DnaA, conserved site | 483 | 502 | - |
| CDD | cd00009 | AAA | 203 | 346 | 2.49461E-17 | ||
| Gene3D | G3DSA:1.10.8.60 | 356 | 425 | 2.9E-24 | |||
| CDD | cd06571 | Bac_DnaA_C | IPR013159 | Chromosomal replication initiator, DnaA C-terminal | 434 | 523 | 3.19193E-43 |
| Pfam | PF00308 | Bacterial dnaA protein | IPR013317 | Chromosomal replication initiator protein DnaA | 190 | 407 | 8.5E-86 |
| Gene3D | G3DSA:1.10.1750.10 | IPR010921 | Trp repressor/replication initiator | 426 | 524 | 2.4E-37 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.