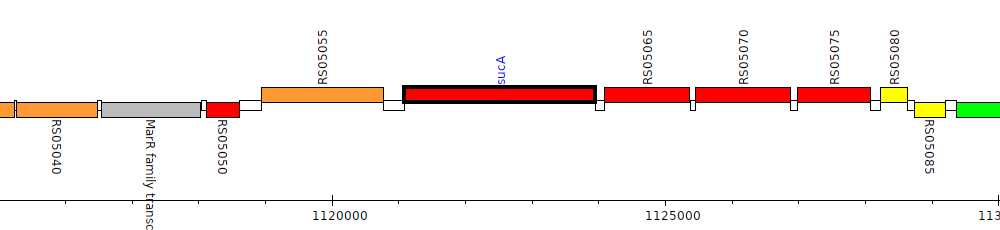
Burkholderia cenocepacia AU 1054, BCEN_RS05060 (sucA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006099 | tricarboxylic acid cycle |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000157
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016624 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, disulfide as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PF00676
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000157
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000157
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030976 | thiamine pyrophosphate binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000157
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52518 | IPR029061 | Thiamin diphosphate-binding fold | 126 | 552 | 6.42E-90 | |
| Gene3D | G3DSA:3.40.50.970 | 172 | 535 | 4.3E-147 | |||
| Gene3D | G3DSA:3.40.50.11610 | IPR042179 | Multifunctional 2-oxoglutarate metabolism enzyme, C-terminal domain superfamily | 734 | 941 | 3.7E-173 | |
| Pfam | PF00676 | Dehydrogenase E1 component | IPR001017 | Dehydrogenase, E1 component | 220 | 524 | 2.7E-50 |
| Gene3D | G3DSA:3.40.50.12470 | 563 | 813 | 3.7E-173 | |||
| CDD | cd02016 | TPP_E1_OGDC_like | 220 | 481 | 6.75422E-167 | ||
| Gene3D | G3DSA:1.10.287.1150 | 89 | 171 | 1.5E-22 | |||
| Pfam | PF02779 | Transketolase, pyrimidine binding domain | IPR005475 | Transketolase-like, pyrimidine-binding domain | 599 | 798 | 6.4E-66 |
| PIRSF | PIRSF000157 | IPR011603 | 2-oxoglutarate dehydrogenase E1 component | 1 | 946 | 0.0 | |
| TIGRFAM | TIGR00239 | 2oxo_dh_E1: oxoglutarate dehydrogenase (succinyl-transferring), E1 component | IPR011603 | 2-oxoglutarate dehydrogenase E1 component | 14 | 943 | 0.0 |
| Pfam | PF16078 | 2-oxoglutarate dehydrogenase N-terminus | IPR032106 | 2-oxoglutarate dehydrogenase E1 component, N-terminal domain | 12 | 49 | 1.1E-17 |
| SMART | SM00861 | IPR005475 | Transketolase-like, pyrimidine-binding domain | 600 | 797 | 1.1E-61 | |
| SUPERFAMILY | SSF52518 | IPR029061 | Thiamin diphosphate-binding fold | 583 | 804 | 5.74E-51 | |
| Pfam | PF16870 | 2-oxoglutarate dehydrogenase C-terminal | IPR031717 | Multifunctional 2-oxoglutarate metabolism enzyme, C-terminal | 802 | 944 | 3.3E-51 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.