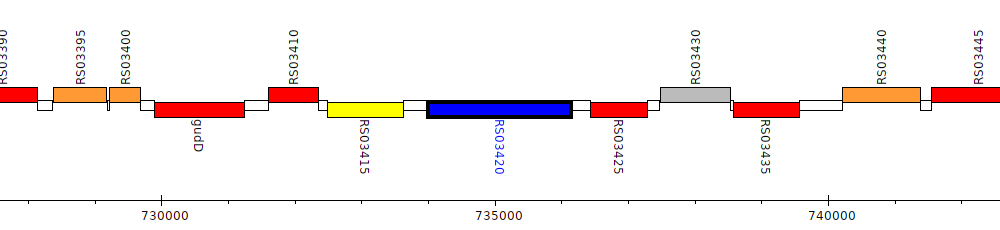
Burkholderia cenocepacia AU 1054, BCEN_RS03420
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004629 | phospholipase C activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF05506
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.720.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004629 | phospholipase C activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF05506
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016788 | hydrolase activity, acting on ester bonds |
Inferred from Sequence Model
Term mapped from: InterPro:PF04185
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.720.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016042 | lipid catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF05506
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0034480 | phosphatidylcholine phospholipase C activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03396
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016042 | lipid catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF05506
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016788 | hydrolase activity, acting on ester bonds |
Inferred from Sequence Model
Term mapped from: InterPro:PF04185
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0034480 | phosphatidylcholine phospholipase C activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03396
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00562 | Inositol phosphate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | plasmalogen biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00564 | Glycerophospholipid metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | 2-arachidonoylglycerol biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00565 | Ether lipid metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | plasmalogen degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd16014 | PLC | 47 | 432 | 3.31096E-151 | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 688 | 714 | - | ||
| Pfam | PF04185 | Phosphoesterase family | IPR007312 | Phosphoesterase | 47 | 430 | 2.8E-126 |
| TIGRFAM | TIGR03396 | PC_PLC: phospholipase C, phosphocholine-specific | IPR017767 | Bacterial phospholipase C, phosphocholine-specific | 5 | 696 | 3.8E-279 |
| ProSiteProfiles | PS51318 | Twin arginine translocation (Tat) signal profile. | IPR006311 | Twin-arginine translocation pathway, signal sequence | 1 | 34 | 9.253 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 699 | 714 | - | ||
| Pfam | PF05506 | Domain of unknown function (DUF756) | IPR008475 | Bacterial phospholipase C, C-terminal domain | 503 | 589 | 2.0E-24 |
| Gene3D | G3DSA:3.40.720.10 | IPR017850 | Alkaline-phosphatase-like, core domain superfamily | 37 | 190 | 5.8E-29 | |
| Pfam | PF05506 | Domain of unknown function (DUF756) | IPR008475 | Bacterial phospholipase C, C-terminal domain | 602 | 683 | 9.3E-20 |
| Gene3D | G3DSA:3.40.720.10 | IPR017850 | Alkaline-phosphatase-like, core domain superfamily | 278 | 483 | 1.8E-50 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.