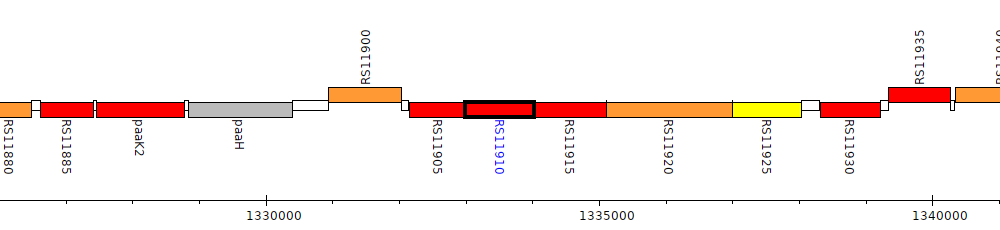
Burkholderia cenocepacia K56-2Valvano, BURCENK562V_RS11910
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0051287 | NAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00389
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PF00389
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00389
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd12167 | 2-Hacid_dh_8 | 3 | 326 | 4.89322E-89 | ||
| Gene3D | G3DSA:3.40.50.720 | 104 | 292 | 8.1E-76 | |||
| Gene3D | G3DSA:3.40.50.720 | 17 | 312 | 8.1E-76 | |||
| Pfam | PF00389 | D-isomer specific 2-hydroxyacid dehydrogenase, catalytic domain | IPR006139 | D-isomer specific 2-hydroxyacid dehydrogenase, catalytic domain | 23 | 318 | 3.2E-14 |
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 109 | 286 | 5.44E-45 | |
| SUPERFAMILY | SSF52283 | 16 | 128 | 1.11E-19 | |||
| ProSitePatterns | PS00671 | D-isomer specific 2-hydroxyacid dehydrogenases signature 3. | IPR029753 | D-isomer specific 2-hydroxyacid dehydrogenase, NAD-binding domain conserved site | 225 | 241 | - |
| Pfam | PF02826 | D-isomer specific 2-hydroxyacid dehydrogenase, NAD binding domain | IPR006140 | D-isomer specific 2-hydroxyacid dehydrogenase, NAD-binding domain | 116 | 286 | 2.5E-47 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.