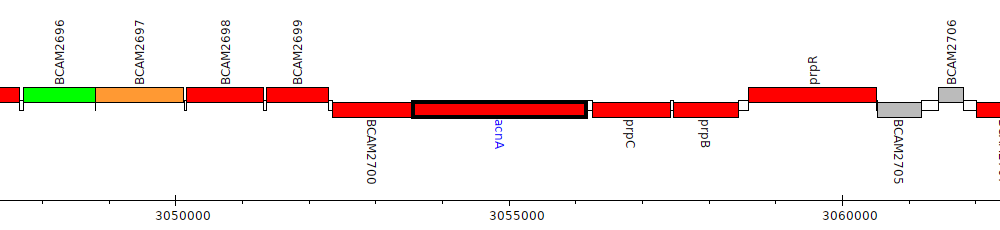
Burkholderia cenocepacia J2315, BCAM2701 (acnA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0019679 | propionate metabolic process, methylcitrate cycle |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02333
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00630 | Glyoxylate and dicarboxylate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01210 | 2-Oxocarboxylic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00720 | Carbon fixation pathways in prokaryotes | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00020 | Citrate cycle (TCA cycle) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00020 | Citrate cycle (TCA cycle) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00630 | Glyoxylate and dicarboxylate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00290 | Valine, leucine and isoleucine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00330 | Aconitase family (aconitate hydratase) | IPR001030 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha domain | 65 | 533 | 3.4E-147 |
| PRINTS | PR00415 | Aconitase family signature | IPR001030 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha domain | 138 | 151 | 3.7E-28 |
| PRINTS | PR00415 | Aconitase family signature | IPR001030 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha domain | 191 | 204 | 3.7E-28 |
| SUPERFAMILY | SSF53732 | IPR036008 | Aconitase, iron-sulfur domain | 15 | 589 | 8.5E-188 | |
| PRINTS | PR00415 | Aconitase family signature | IPR001030 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha domain | 360 | 374 | 3.7E-28 |
| PRINTS | PR00415 | Aconitase family signature | IPR001030 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha domain | 281 | 294 | 3.7E-28 |
| PRINTS | PR00415 | Aconitase family signature | IPR001030 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha domain | 267 | 280 | 3.7E-28 |
| Gene3D | G3DSA:3.20.19.10 | IPR015928 | Aconitase/3-isopropylmalate dehydratase, swivel | 620 | 858 | 5.8E-84 | |
| PRINTS | PR00415 | Aconitase family signature | IPR001030 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha domain | 464 | 477 | 3.7E-28 |
| Gene3D | G3DSA:3.30.499.20 | 9 | 360 | 2.1E-131 | |||
| TIGRFAM | TIGR02333 | 2met_isocit_dHY: 2-methylisocitrate dehydratase, Fe/S-dependent | IPR012708 | 2-methylisocitrate dehydratase AcnD, Fe/S-dependent | 2 | 859 | 0.0 |
| Pfam | PF00694 | Aconitase C-terminal domain | IPR000573 | Aconitase A/isopropylmalate dehydratase small subunit, swivel domain | 657 | 787 | 8.4E-28 |
| Gene3D | G3DSA:1.10.1440.20 | 562 | 619 | 6.0E-9 | |||
| SUPERFAMILY | SSF52016 | 598 | 857 | 2.35E-67 | |||
| PRINTS | PR00415 | Aconitase family signature | IPR001030 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha domain | 205 | 220 | 3.7E-28 |
| Gene3D | G3DSA:3.30.499.10 | IPR015931 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha, subdomain 1/3 | 361 | 561 | 3.0E-76 | |
| PRINTS | PR00415 | Aconitase family signature | IPR001030 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha domain | 162 | 170 | 3.7E-28 |
| PRINTS | PR00415 | Aconitase family signature | IPR001030 | Aconitase/3-isopropylmalate dehydratase large subunit, alpha/beta/alpha domain | 402 | 413 | 3.7E-28 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.