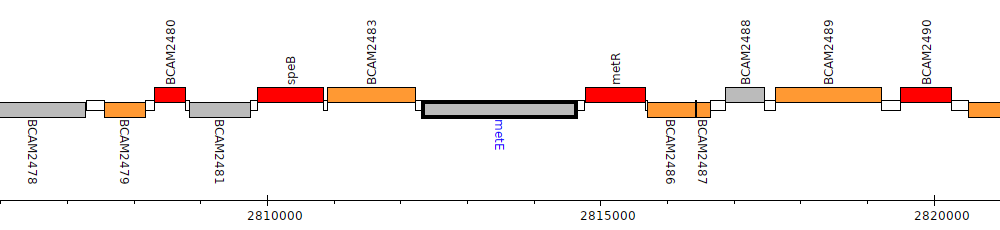
Burkholderia cenocepacia J2315, BCAM2484 (metE)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009086 | methionine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003871 | 5-methyltetrahydropteroyltriglutamate-homocysteine S-methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0008652 | cellular amino acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF08267
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009086 | methionine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003871 | 5-methyltetrahydropteroyltriglutamate-homocysteine S-methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0008652 | cellular amino acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF08267
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000382
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | <i>S</i>-adenosyl-L-methionine cycle II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00270 | Cysteine and methionine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-methionine biosynthesis II (plants) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | seleno-amino acid biosynthesis (plants) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00270 | Cysteine and methionine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00450 | Selenocompound metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00450 | Selenocompound metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | <i>S</i>-adenosyl-L-methionine cycle I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF51726 | IPR038071 | UROD/MetE-like superfamily | 400 | 762 | 4.32E-118 | |
| Hamap | MF_00172 | 5-methyltetrahydropteroyltriglutamate--homocysteine methyltransferase [metE]. | IPR006276 | Cobalamin-independent methionine synthase | 2 | 762 | 37.405 |
| Gene3D | G3DSA:3.20.20.210 | IPR038071 | UROD/MetE-like superfamily | 393 | 760 | 0.0 | |
| Gene3D | G3DSA:3.20.20.210 | IPR038071 | UROD/MetE-like superfamily | 6 | 528 | 0.0 | |
| SUPERFAMILY | SSF51726 | IPR038071 | UROD/MetE-like superfamily | 4 | 398 | 4.97E-109 | |
| Pfam | PF01717 | Cobalamin-independent synthase, Catalytic domain | IPR002629 | Cobalamin-independent methionine synthase MetE, C-terminal/archaeal | 435 | 757 | 2.6E-155 |
| TIGRFAM | TIGR01371 | met_syn_B12ind: 5-methyltetrahydropteroyltriglutamate--homocysteine S-methyltransferase | IPR006276 | Cobalamin-independent methionine synthase | 7 | 762 | 0.0 |
| CDD | cd03312 | CIMS_N_terminal_like | 3 | 368 | 0.0 | ||
| PIRSF | PIRSF000382 | IPR006276 | Cobalamin-independent methionine synthase | 1 | 764 | 0.0 | |
| CDD | cd03311 | CIMS_C_terminal_like | 436 | 757 | 1.48399E-146 | ||
| Pfam | PF08267 | Cobalamin-independent synthase, N-terminal domain | IPR013215 | Cobalamin-independent methionine synthase MetE, N-terminal | 4 | 319 | 2.3E-113 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.