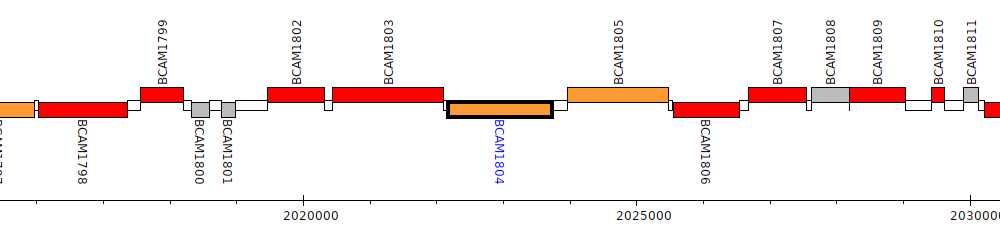
Burkholderia cenocepacia J2315, BCAM1804
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006935 | chemotaxis |
Inferred from Sequence Model
Term mapped from: InterPro:PF02203
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004888 | transmembrane signaling receptor activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PS50111
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:SM00304
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:SM00304
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj02030 | Bacterial chemotaxis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj02020 | Two-component system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSiteProfiles | PS50885 | HAMP domain profile. | IPR003660 | HAMP domain | 217 | 269 | 9.513 |
| CDD | cd06225 | HAMP | IPR003660 | HAMP domain | 219 | 266 | 1.12376E-5 |
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 291 | 320 | 4.5E-59 |
| ProSiteProfiles | PS50111 | Bacterial chemotaxis sensory transducers domain profile. | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 274 | 503 | 49.475 |
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 397 | 426 | 4.5E-59 |
| ProSitePatterns | PS00538 | Bacterial chemotaxis sensory transducers signature. | IPR004091 | Chemotaxis methyl-accepting receptor, methyl-accepting site | 295 | 317 | - |
| Gene3D | G3DSA:1.10.287.950 | 217 | 519 | 1.8E-89 | |||
| Pfam | PF02203 | Tar ligand binding domain homologue | IPR003122 | Chemotaxis methyl-accepting receptor Tar-related, ligand-binding | 7 | 178 | 3.7E-49 |
| CDD | cd11386 | MCP_signal | 297 | 488 | 1.94184E-55 | ||
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 445 | 474 | 4.5E-59 |
| Coils | Coil | 293 | 313 | - | |||
| Pfam | PF00015 | Methyl-accepting chemotaxis protein (MCP) signalling domain | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 330 | 485 | 4.2E-51 |
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 10 | 35 | 4.5E-59 |
| SUPERFAMILY | SSF47170 | IPR035440 | Methyl-accepting chemotaxis protein, four helix bundle domain superfamily | 31 | 151 | 1.96E-9 | |
| SUPERFAMILY | SSF58104 | 237 | 518 | 2.49E-71 | |||
| SMART | SM00304 | IPR003660 | HAMP domain | 217 | 269 | 1.0E-7 | |
| Pfam | PF00672 | HAMP domain | IPR003660 | HAMP domain | 218 | 265 | 4.5E-7 |
| Coils | Coil | 474 | 512 | - | |||
| CDD | cd00181 | Tar_Tsr_LBD | IPR003122 | Chemotaxis methyl-accepting receptor Tar-related, ligand-binding | 50 | 175 | 1.20162E-13 |
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 368 | 395 | 4.5E-59 |
| SMART | SM00283 | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 270 | 517 | 1.9E-84 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.