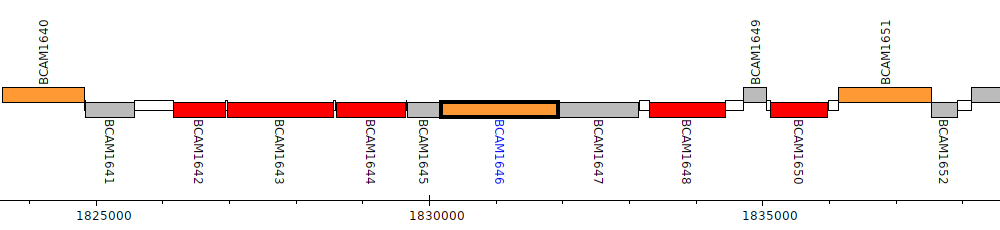
Burkholderia cenocepacia J2315, BCAM1646
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00984 | Steroid degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF56425 | IPR027477 | Succinate dehydrogenase/fumarate reductase flavoprotein, catalytic domain superfamily | 341 | 507 | 2.88E-30 | |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 523 | 538 | 2.3E-7 | ||
| SUPERFAMILY | SSF51905 | IPR036188 | FAD/NAD(P)-binding domain superfamily | 10 | 567 | 2.02E-41 | |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | 12 | 34 | 2.3E-7 | ||
| Pfam | PF00890 | FAD binding domain | IPR003953 | FAD-dependent oxidoreductase 2, FAD binding domain | 12 | 548 | 2.4E-86 |
| ProSiteProfiles | PS51257 | Prokaryotic membrane lipoprotein lipid attachment site profile. | 1 | 27 | 5.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.