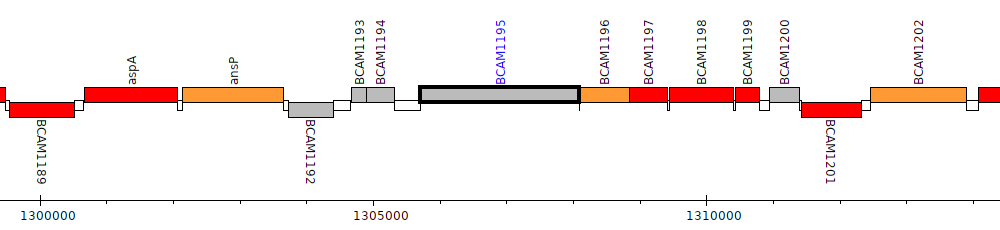
Burkholderia cenocepacia J2315, BCAM1195
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:SM00304
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence Model
Term mapped from: InterPro:cd00088
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:SM00304
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.20.120.160 | IPR036641 | HPT domain superfamily | 417 | 547 | 2.1E-11 | |
| SMART | SM00086 | IPR001610 | PAC motif | 352 | 394 | 4.3E-4 | |
| CDD | cd16916 | HATPase_CheA-like | 595 | 745 | 4.47166E-47 | ||
| Coils | Coil | 151 | 171 | - | |||
| Pfam | PF00672 | HAMP domain | IPR003660 | HAMP domain | 204 | 257 | 4.3E-8 |
| Coils | Coil | 470 | 490 | - | |||
| SMART | SM00304 | IPR003660 | HAMP domain | 207 | 260 | 2.6E-7 | |
| Pfam | PF02518 | Histidine kinase-, DNA gyrase B-, and HSP90-like ATPase | IPR003594 | Histidine kinase/HSP90-like ATPase | 624 | 747 | 3.6E-7 |
| SUPERFAMILY | SSF55785 | IPR035965 | PAS domain superfamily | 347 | 396 | 9.9E-6 | |
| Gene3D | G3DSA:3.30.565.10 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 581 | 756 | 7.5E-32 | |
| SUPERFAMILY | SSF47226 | IPR036641 | HPT domain superfamily | 419 | 515 | 1.57E-13 | |
| ProSiteProfiles | PS50885 | HAMP domain profile. | IPR003660 | HAMP domain | 207 | 260 | 10.173 |
| Gene3D | G3DSA:3.30.450.20 | 268 | 410 | 1.9E-6 | |||
| Coils | Coil | 245 | 272 | - | |||
| Coils | Coil | 76 | 96 | - | |||
| ProSiteProfiles | PS50894 | Histidine-containing phosphotransfer (HPt) domain profile. | IPR008207 | Signal transduction histidine kinase, phosphotransfer (Hpt) domain | 414 | 522 | 14.781 |
| SMART | SM00387 | IPR003594 | Histidine kinase/HSP90-like ATPase | 622 | 771 | 2.3E-14 | |
| CDD | cd00088 | HPT | IPR008207 | Signal transduction histidine kinase, phosphotransfer (Hpt) domain | 432 | 513 | 6.13942E-7 |
| SUPERFAMILY | SSF158472 | 207 | 257 | 1.1E-6 | |||
| SUPERFAMILY | SSF55874 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 605 | 747 | 2.23E-25 | |
| ProSiteProfiles | PS50113 | PAC domain profile. | IPR000700 | PAS-associated, C-terminal | 351 | 403 | 10.189 |
| CDD | cd06225 | HAMP | IPR003660 | HAMP domain | 209 | 257 | 2.18261E-6 |
| Gene3D | G3DSA:1.20.1480.50 | 166 | 267 | 5.0E-9 | |||
| Pfam | PF12729 | Four helix bundle sensory module for signal transduction | IPR024478 | Chemotaxis methyl-accepting receptor HlyB-like, 4HB MCP domain | 1 | 161 | 1.5E-11 |
| Pfam | PF01627 | Hpt domain | IPR008207 | Signal transduction histidine kinase, phosphotransfer (Hpt) domain | 447 | 506 | 8.5E-7 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.