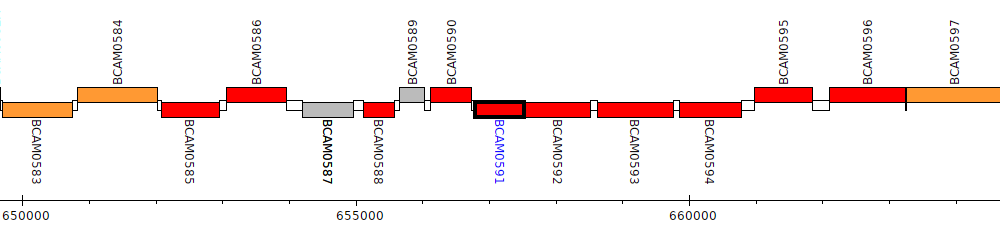
Burkholderia cenocepacia J2315, BCAM0591
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01509
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF56784 | IPR036412 | HAD-like superfamily | 7 | 233 | 1.3E-39 | |
| SFLD | SFLDS00003 | Haloacid Dehalogenase | 6 | 226 | 9.2E-24 | ||
| SFLD | SFLDG01129 | C1.5: HAD, Beta-PGM, Phosphatase Like | 6 | 226 | 9.2E-24 | ||
| PRINTS | PR00413 | Haloacid dehalogenase/epoxide hydrolase family signature | IPR006439 | HAD hydrolase, subfamily IA | 162 | 182 | 1.5E-5 |
| Pfam | PF00702 | haloacid dehalogenase-like hydrolase | 7 | 195 | 3.2E-12 | ||
| TIGRFAM | TIGR01549 | HAD-SF-IA-v1: HAD hydrolase, family IA, variant 1 | IPR006439 | HAD hydrolase, subfamily IA | 88 | 195 | 2.9E-10 |
| TIGRFAM | TIGR01509 | HAD-SF-IA-v3: HAD hydrolase, family IA, variant 3 | IPR006439 | HAD hydrolase, subfamily IA | 97 | 201 | 9.4E-9 |
| PRINTS | PR00413 | Haloacid dehalogenase/epoxide hydrolase family signature | IPR006439 | HAD hydrolase, subfamily IA | 144 | 160 | 1.5E-5 |
| Gene3D | G3DSA:3.40.50.1000 | IPR023214 | HAD superfamily | 8 | 216 | 2.0E-54 | |
| Gene3D | G3DSA:1.20.120.1600 | 20 | 104 | 2.0E-54 | |||
| PRINTS | PR00413 | Haloacid dehalogenase/epoxide hydrolase family signature | IPR006439 | HAD hydrolase, subfamily IA | 6 | 17 | 1.5E-5 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.