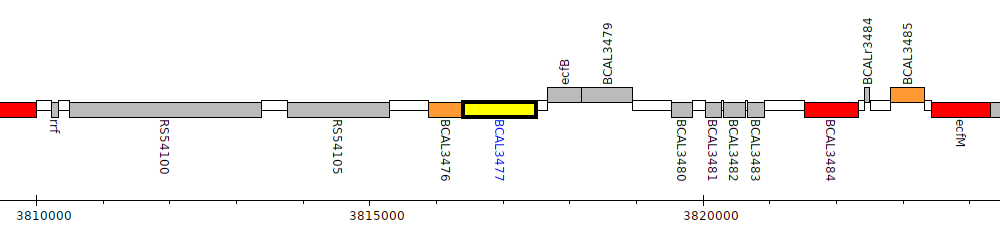
Burkholderia cenocepacia J2315, BCAL3477
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56634
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006979 | response to oxidative stress |
Inferred from Sequence Model
Term mapped from: InterPro:PR00067
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004096 | catalase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00067
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0020037 | heme binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56634
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00380 | Tryptophan metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00380 | Tryptophan metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00630 | Glyoxylate and dicarboxylate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00630 | Glyoxylate and dicarboxylate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.20.1280.120 | 51 | 343 | 3.0E-108 | |||
| PRINTS | PR00067 | Catalase signature | IPR018028 | Catalase, mono-functional, haem-containing | 96 | 114 | 8.2E-6 |
| Gene3D | G3DSA:2.40.180.20 | 72 | 329 | 3.0E-108 | |||
| ProSiteProfiles | PS51402 | catalase family profile. | IPR018028 | Catalase, mono-functional, haem-containing | 1 | 364 | 16.664 |
| PRINTS | PR00067 | Catalase signature | IPR018028 | Catalase, mono-functional, haem-containing | 322 | 348 | 8.2E-6 |
| Pfam | PF00199 | Catalase | IPR011614 | Catalase core domain | 65 | 348 | 2.7E-32 |
| PRINTS | PR00067 | Catalase signature | IPR018028 | Catalase, mono-functional, haem-containing | 59 | 81 | 8.2E-6 |
| SMART | SM01060 | IPR011614 | Catalase core domain | 21 | 364 | 1.6E-9 | |
| CDD | cd08153 | srpA_like | IPR024168 | Catalase, SrpA-type, predicted | 52 | 347 | 1.97629E-159 |
| SUPERFAMILY | SSF56634 | IPR020835 | Catalase superfamily | 36 | 348 | 3.27E-96 | |
| PIRSF | PIRSF000296 | IPR024168 | Catalase, SrpA-type, predicted | 18 | 350 | 7.7E-122 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.