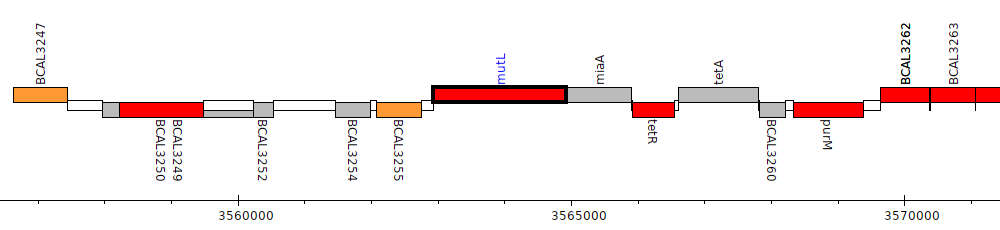
Burkholderia cenocepacia J2315, BCAL3256 (mutL)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00585
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006298 | mismatch repair |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00585
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030983 | mismatched DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00585
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj03430 | Mismatch repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF08676 | MutL C terminal dimerisation domain | IPR014790 | MutL, C-terminal, dimerisation | 474 | 621 | 8.5E-46 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 375 | 405 | - | ||
| SUPERFAMILY | SSF54211 | IPR020568 | Ribosomal protein S5 domain 2-type fold | 207 | 342 | 1.4E-35 | |
| Gene3D | G3DSA:2.30.42.20 | IPR042120 | MutL, C-terminal domain, dimerisation subdomain | 480 | 658 | 1.3E-57 | |
| SUPERFAMILY | SSF55874 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 23 | 223 | 1.83E-52 | |
| CDD | cd03482 | MutL_Trans_MutL | 221 | 342 | 1.15882E-67 | ||
| Gene3D | G3DSA:3.30.230.10 | IPR014721 | Ribosomal protein S5 domain 2-type fold, subgroup | 230 | 341 | 1.6E-36 | |
| Pfam | PF01119 | DNA mismatch repair protein, C-terminal domain | IPR013507 | DNA mismatch repair protein, S5 domain 2-like | 225 | 341 | 3.2E-37 |
| CDD | cd16926 | HATPase_MutL-MLH-PMS-like | 30 | 210 | 6.57111E-99 | ||
| SMART | SM01340 | IPR013507 | DNA mismatch repair protein, S5 domain 2-like | 224 | 342 | 8.3E-56 | |
| TIGRFAM | TIGR00585 | mutl: DNA mismatch repair protein MutL | IPR002099 | DNA mismatch repair protein family, N-terminal | 22 | 322 | 1.9E-91 |
| ProSitePatterns | PS00058 | DNA mismatch repair proteins mutL / hexB / PMS1 signature. | IPR014762 | DNA mismatch repair, conserved site | 113 | 119 | - |
| Hamap | MF_00149 | DNA mismatch repair protein MutL [mutL]. | IPR020667 | DNA mismatch repair protein, MutL | 21 | 661 | 27.942 |
| SUPERFAMILY | SSF118116 | IPR037198 | MutL, C-terminal domain superfamily | 474 | 663 | 5.49E-58 | |
| SMART | SM00853 | IPR014790 | MutL, C-terminal, dimerisation | 478 | 621 | 2.3E-51 | |
| Pfam | PF13589 | Histidine kinase-, DNA gyrase B-, and HSP90-like ATPase | 44 | 145 | 8.9E-12 | ||
| Gene3D | G3DSA:3.30.565.10 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 22 | 229 | 1.6E-64 | |
| Gene3D | G3DSA:3.30.1370.100 | IPR042121 | MutL, C-terminal domain, regulatory subdomain | 521 | 616 | 1.3E-57 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.