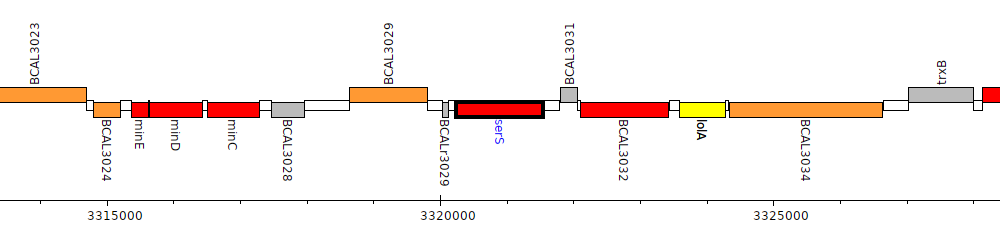
Burkholderia cenocepacia J2315, BCAL3030 (serS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006418 | tRNA aminoacylation for protein translation |
Inferred from Sequence Model
Term mapped from: InterPro:PF00587
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00587
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006434 | seryl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00414
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004828 | serine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00414
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00414
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00414
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00970 | Aminoacyl-tRNA biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00970 | Aminoacyl-tRNA biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-selenocysteine biosynthesis II (archaea and eukaryotes) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00981 | Seryl-tRNA synthetase signature | IPR002317 | Serine-tRNA ligase, type1 | 268 | 280 | 3.2E-32 |
| Hamap | MF_00176 | Serine--tRNA ligase [serS]. | IPR002317 | Serine-tRNA ligase, type1 | 1 | 425 | 39.81 |
| CDD | cd00770 | SerRS_core | IPR033729 | Serine-tRNA ligase catalytic core domain | 119 | 420 | 1.88965E-179 |
| PRINTS | PR00981 | Seryl-tRNA synthetase signature | IPR002317 | Serine-tRNA ligase, type1 | 280 | 293 | 3.2E-32 |
| TIGRFAM | TIGR00414 | serS: serine--tRNA ligase | IPR002317 | Serine-tRNA ligase, type1 | 1 | 420 | 3.7E-163 |
| Gene3D | G3DSA:1.10.287.40 | IPR042103 | Serine-tRNA synthetase, type1, N-terminal domain superfamily | 1 | 104 | 4.9E-24 | |
| PRINTS | PR00981 | Seryl-tRNA synthetase signature | IPR002317 | Serine-tRNA ligase, type1 | 355 | 371 | 3.2E-32 |
| SUPERFAMILY | SSF46589 | IPR010978 | Class I and II aminoacyl-tRNA synthetase, tRNA-binding arm | 1 | 112 | 2.3E-30 | |
| PRINTS | PR00981 | Seryl-tRNA synthetase signature | IPR002317 | Serine-tRNA ligase, type1 | 320 | 333 | 3.2E-32 |
| Gene3D | G3DSA:3.30.930.10 | 105 | 431 | 1.8E-117 | |||
| Pfam | PF02403 | Seryl-tRNA synthetase N-terminal domain | IPR015866 | Serine-tRNA synthetase, type1, N-terminal | 1 | 106 | 9.4E-32 |
| SUPERFAMILY | SSF55681 | 116 | 421 | 1.82E-99 | |||
| PRINTS | PR00981 | Seryl-tRNA synthetase signature | IPR002317 | Serine-tRNA ligase, type1 | 337 | 353 | 3.2E-32 |
| PIRSF | PIRSF001529 | IPR002317 | Serine-tRNA ligase, type1 | 1 | 429 | 3.0E-182 | |
| ProSiteProfiles | PS50862 | Aminoacyl-transfer RNA synthetases class-II family profile. | IPR006195 | Aminoacyl-tRNA synthetase, class II | 170 | 413 | 25.19 |
| Pfam | PF00587 | tRNA synthetase class II core domain (G, H, P, S and T) | IPR002314 | Aminoacyl-tRNA synthetase, class II (G/ P/ S/T) | 224 | 403 | 4.4E-33 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.