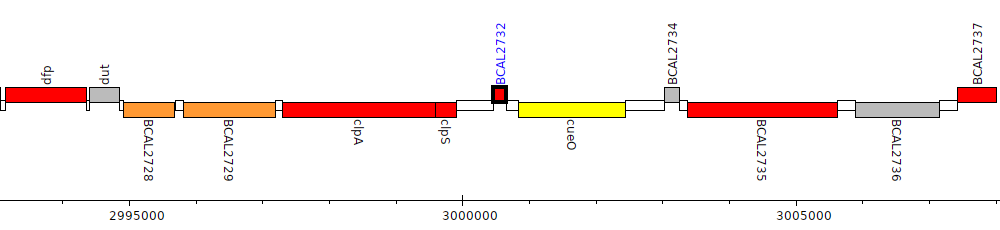
Burkholderia cenocepacia J2315, BCAL2732
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF002599
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00050 | Cold shock protein signature | IPR002059 | Cold-shock protein, DNA-binding | 40 | 58 | 6.0E-18 |
| PRINTS | PR00050 | Cold shock protein signature | IPR002059 | Cold-shock protein, DNA-binding | 4 | 19 | 6.0E-18 |
| ProSitePatterns | PS00352 | Cold-shock (CSD) domain signature. | IPR019844 | Cold-shock (CSD) domain | 15 | 34 | - |
| Gene3D | G3DSA:2.40.50.140 | 1 | 67 | 4.3E-31 | |||
| Pfam | PF00313 | 'Cold-shock' DNA-binding domain | IPR002059 | Cold-shock protein, DNA-binding | 3 | 66 | 4.1E-31 |
| CDD | cd04458 | CSP_CDS | IPR002059 | Cold-shock protein, DNA-binding | 3 | 65 | 2.1771E-32 |
| SMART | SM00357 | IPR011129 | Cold shock domain | 3 | 67 | 1.0E-31 | |
| PIRSF | PIRSF002599 | IPR012156 | Cold shock, CspA | 1 | 67 | 2.3E-26 | |
| PRINTS | PR00050 | Cold shock protein signature | IPR002059 | Cold-shock protein, DNA-binding | 25 | 34 | 6.0E-18 |
| ProSiteProfiles | PS51857 | Cold-shock (CSD) domain profile. | 1 | 66 | 38.06 | ||
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 1 | 66 | 2.36E-26 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.