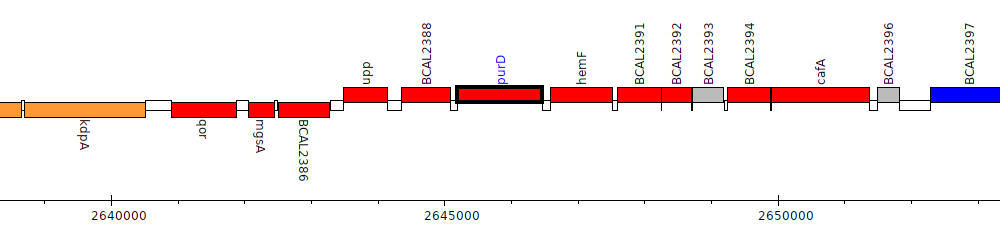
Burkholderia cenocepacia J2315, BCAL2389 (purD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004637 | phosphoribosylamine-glycine ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02843
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50975
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.30.1490.20
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009113 | purine nucleobase biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02843
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 5-aminoimidazole ribonucleotide biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 5-aminoimidazole ribonucleotide biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00230 | Purine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | superpathway of 5-aminoimidazole ribonucleotide biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM01209 | 101 | 294 | 3.5E-138 | |||
| Gene3D | G3DSA:3.30.1490.20 | IPR013815 | ATP-grasp fold, subdomain 1 | 119 | 188 | 1.2E-27 | |
| SUPERFAMILY | SSF52440 | IPR016185 | Pre-ATP-grasp domain superfamily | 1 | 101 | 1.44E-34 | |
| Pfam | PF02844 | Phosphoribosylglycinamide synthetase, N domain | IPR020562 | Phosphoribosylglycinamide synthetase, N-terminal | 1 | 100 | 1.5E-34 |
| SUPERFAMILY | SSF56059 | 102 | 324 | 1.34E-64 | |||
| Gene3D | G3DSA:3.40.50.20 | 1 | 93 | 5.4E-35 | |||
| Pfam | PF02843 | Phosphoribosylglycinamide synthetase, C domain | IPR020560 | Phosphoribosylglycinamide synthetase, C-domain | 329 | 419 | 2.6E-30 |
| SMART | SM01210 | IPR020560 | Phosphoribosylglycinamide synthetase, C-domain | 328 | 421 | 2.4E-52 | |
| Hamap | MF_00138 | Phosphoribosylamine--glycine ligase [purD]. | IPR000115 | Phosphoribosylglycinamide synthetase | 1 | 421 | 43.217 |
| ProSiteProfiles | PS50975 | ATP-grasp fold profile. | IPR011761 | ATP-grasp fold | 107 | 314 | 42.415 |
| SUPERFAMILY | SSF51246 | IPR011054 | Rudiment single hybrid motif | 327 | 421 | 6.43E-32 | |
| Gene3D | G3DSA:3.30.470.20 | 189 | 327 | 7.2E-57 | |||
| Gene3D | G3DSA:3.90.600.10 | IPR037123 | Phosphoribosylglycinamide synthetase, C-domain superfamily | 328 | 426 | 6.0E-34 | |
| TIGRFAM | TIGR00877 | purD: phosphoribosylamine--glycine ligase | IPR000115 | Phosphoribosylglycinamide synthetase | 1 | 421 | 4.1E-174 |
| Pfam | PF01071 | Phosphoribosylglycinamide synthetase, ATP-grasp (A) domain | IPR020561 | Phosphoribosylglycinamide synthetase, ATP-grasp (A) domain | 101 | 294 | 9.7E-78 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.