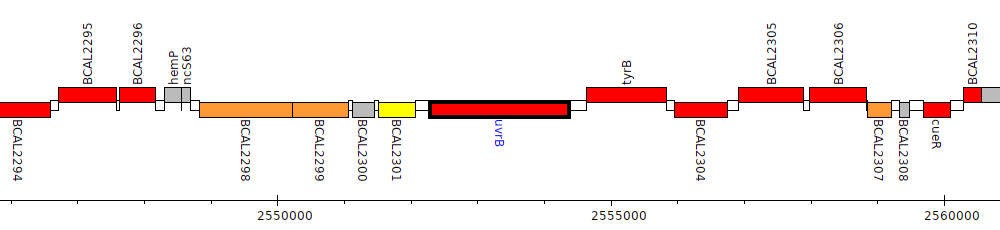
Burkholderia cenocepacia J2315, BCAL2302 (uvrB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02151
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006289 | nucleotide-excision repair |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00204
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00204
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00204
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF04851
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00204
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009380 | excinuclease repair complex |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00204
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj03420 | Nucleotide excision repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00271 | Helicase conserved C-terminal domain | IPR001650 | Helicase, C-terminal | 457 | 563 | 3.7E-18 |
| Coils | Coil | 277 | 297 | - | |||
| Pfam | PF02151 | UvrB/uvrC motif | IPR001943 | UVR domain | 648 | 681 | 3.5E-7 |
| Pfam | PF12344 | Ultra-violet resistance protein B | IPR024759 | UvrB, YAD/RRR-motif-containing domain | 571 | 613 | 2.2E-21 |
| Gene3D | G3DSA:3.40.50.300 | 434 | 615 | 4.6E-52 | |||
| TIGRFAM | TIGR00631 | uvrb: excinuclease ABC subunit B | IPR004807 | UvrABC system, subunit B | 25 | 679 | 0.0 |
| ProSiteProfiles | PS51194 | Superfamilies 1 and 2 helicase C-terminal domain profile. | IPR001650 | Helicase, C-terminal | 450 | 616 | 21.404 |
| Pfam | PF17757 | UvrB interaction domain | IPR041471 | UvrB, interaction domain | 179 | 269 | 2.7E-32 |
| Coils | Coil | 622 | 670 | - | |||
| Pfam | PF04851 | Type III restriction enzyme, res subunit | IPR006935 | Helicase/UvrB, N-terminal | 57 | 158 | 2.5E-8 |
| Gene3D | G3DSA:4.10.860.10 | 630 | 687 | 9.8E-18 | |||
| CDD | cd17916 | DEXHc_UvrB | 25 | 433 | 0.0 | ||
| SMART | SM00487 | IPR014001 | Helicase superfamily 1/2, ATP-binding domain | 29 | 445 | 1.5E-28 | |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 57 | 608 | 6.52E-37 | |
| SUPERFAMILY | SSF46600 | IPR036876 | UVR domain superfamily | 635 | 683 | 2.48E-11 | |
| ProSiteProfiles | PS50151 | UVR domain profile. | IPR001943 | UVR domain | 647 | 682 | 11.884 |
| Gene3D | G3DSA:3.40.50.300 | 288 | 433 | 3.9E-49 | |||
| Gene3D | G3DSA:3.40.50.300 | 24 | 250 | 6.5E-71 | |||
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 24 | 434 | 6.38E-133 | |
| Hamap | MF_00204 | UvrABC system protein B [uvrB]. | IPR004807 | UvrABC system, subunit B | 23 | 682 | 43.223 |
| CDD | cd18790 | SF2_C_UvrB | 439 | 609 | 1.44683E-112 | ||
| SMART | SM00490 | IPR001650 | Helicase, C-terminal | 479 | 565 | 1.1E-18 | |
| ProSiteProfiles | PS51192 | Superfamilies 1 and 2 helicase ATP-binding type-1 domain profile. | IPR014001 | Helicase superfamily 1/2, ATP-binding domain | 46 | 180 | 14.231 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.