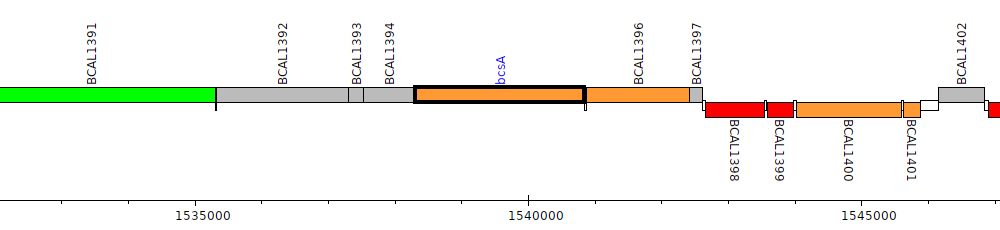
Burkholderia cenocepacia J2315, BCAL1395 (bcsA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016760 | cellulose synthase (UDP-forming) activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF03552
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03030
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0035438 | cyclic-di-GMP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03030
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0030244 | cellulose biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF03552
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016759 | cellulose synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03030
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006011 | UDP-glucose metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03030
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00500 | Starch and sucrose metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | cellulose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00500 | Starch and sucrose metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR01439 | Cellulose synthase subunit A signature | IPR003919 | Cellulose synthase, subunit A | 665 | 686 | 2.1E-44 |
| PRINTS | PR01439 | Cellulose synthase subunit A signature | IPR003919 | Cellulose synthase, subunit A | 197 | 217 | 2.1E-44 |
| PRINTS | PR01439 | Cellulose synthase subunit A signature | IPR003919 | Cellulose synthase, subunit A | 551 | 570 | 2.1E-44 |
| SUPERFAMILY | SSF141371 | 694 | 783 | 1.1E-7 | |||
| PRINTS | PR01439 | Cellulose synthase subunit A signature | IPR003919 | Cellulose synthase, subunit A | 280 | 299 | 2.1E-44 |
| PRINTS | PR01439 | Cellulose synthase subunit A signature | IPR003919 | Cellulose synthase, subunit A | 591 | 610 | 2.1E-44 |
| Gene3D | G3DSA:3.90.550.10 | IPR029044 | Nucleotide-diphospho-sugar transferases | 261 | 481 | 5.2E-81 | |
| PRINTS | PR01439 | Cellulose synthase subunit A signature | IPR003919 | Cellulose synthase, subunit A | 361 | 385 | 2.1E-44 |
| PRINTS | PR01439 | Cellulose synthase subunit A signature | IPR003919 | Cellulose synthase, subunit A | 225 | 250 | 2.1E-44 |
| SUPERFAMILY | SSF53448 | IPR029044 | Nucleotide-diphospho-sugar transferases | 270 | 556 | 1.55E-52 | |
| PRINTS | PR01439 | Cellulose synthase subunit A signature | IPR003919 | Cellulose synthase, subunit A | 522 | 548 | 2.1E-44 |
| CDD | cd06421 | CESA_CelA_like | 271 | 503 | 4.81151E-111 | ||
| Pfam | PF03552 | Cellulose synthase | IPR005150 | Cellulose synthase | 448 | 577 | 2.6E-10 |
| Pfam | PF07238 | PilZ domain | IPR009875 | PilZ domain | 692 | 788 | 7.3E-12 |
| Pfam | PF00535 | Glycosyl transferase family 2 | IPR001173 | Glycosyltransferase 2-like | 274 | 445 | 2.4E-34 |
| Gene3D | G3DSA:2.40.10.220 | 688 | 798 | 1.3E-16 | |||
| TIGRFAM | TIGR03030 | CelA: cellulose synthase catalytic subunit (UDP-forming) | IPR003919 | Cellulose synthase, subunit A | 149 | 826 | 4.4E-267 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.