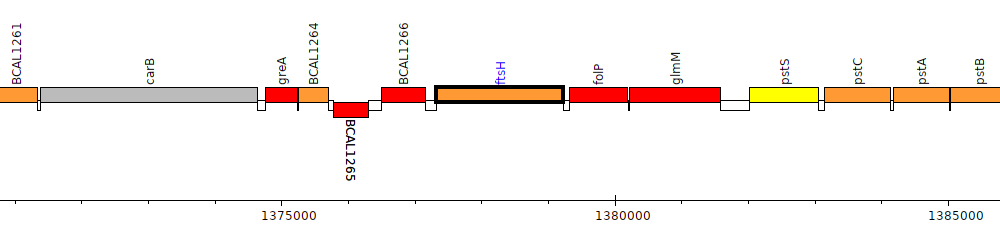
Burkholderia cenocepacia J2315, BCAL1267 (ftsH)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.20.58.760
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF06480
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.20.58.760
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF06480
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004222 | metalloendopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01458
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01458
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00382 | IPR003593 | AAA+ ATPase domain | 186 | 325 | 1.3E-26 | |
| Pfam | PF01434 | Peptidase family M41 | IPR000642 | Peptidase M41 | 404 | 591 | 1.0E-66 |
| Gene3D | G3DSA:1.20.58.760 | IPR037219 | Peptidase M41-like | 399 | 596 | 2.2E-70 | |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 149 | 393 | 7.84E-75 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 589 | 632 | - | ||
| CDD | cd00009 | AAA | 157 | 323 | 2.67089E-32 | ||
| Gene3D | G3DSA:2.40.50.920 | 30 | 103 | 2.5E-9 | |||
| Gene3D | G3DSA:1.10.8.60 | 325 | 398 | 3.8E-30 | |||
| Pfam | PF17862 | AAA+ lid domain | IPR041569 | AAA ATPase, AAA+ lid domain | 345 | 389 | 3.4E-15 |
| SUPERFAMILY | SSF140990 | IPR037219 | Peptidase M41-like | 404 | 601 | 2.09E-69 | |
| Gene3D | G3DSA:3.40.50.300 | 139 | 324 | 3.9E-71 | |||
| Pfam | PF00004 | ATPase family associated with various cellular activities (AAA) | IPR003959 | ATPase, AAA-type, core | 190 | 322 | 4.1E-46 |
| TIGRFAM | TIGR01241 | FtsH_fam: ATP-dependent metallopeptidase HflB | IPR005936 | Peptidase, FtsH | 104 | 592 | 9.6E-230 |
| Pfam | PF06480 | FtsH Extracellular | IPR011546 | Peptidase M41, FtsH extracellular | 11 | 76 | 1.0E-6 |
| ProSitePatterns | PS00674 | AAA-protein family signature. | IPR003960 | ATPase, AAA-type, conserved site | 293 | 311 | - |
| Hamap | MF_01458 | ATP-dependent zinc metalloprotease FtsH [ftsH]. | IPR005936 | Peptidase, FtsH | 4 | 598 | 36.968 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.