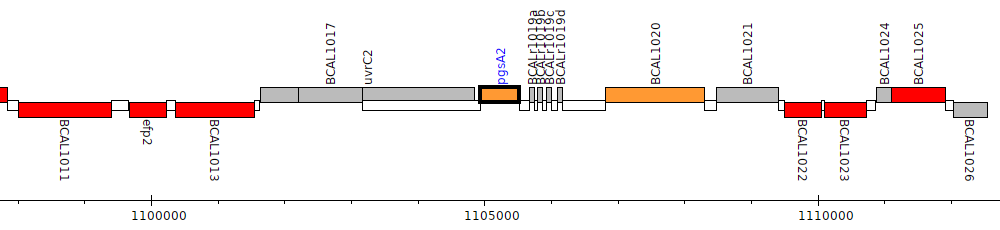
Burkholderia cenocepacia J2315, BCAL1019 (pgsA2)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00560
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0008654 | phospholipid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF01066
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008444 | CDP-diacylglycerol-glycerol-3-phosphate 3-phosphatidyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00560
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016780 | phosphotransferase activity, for other substituted phosphate groups |
Inferred from Sequence Model
Term mapped from: InterPro:PF01066
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF01066
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00564 | Glycerophospholipid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00564 | Glycerophospholipid metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | cardiolipin biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | type I lipoteichoic acid biosynthesis (<i>S. aureus</i>) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | cardiolipin biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF000847 | IPR004570 | CDP-diacylglycerol--glycerol-3-phosphate 3-phosphatidyltransferase | 1 | 195 | 6.7E-48 | |
| Pfam | PF01066 | CDP-alcohol phosphatidyltransferase | IPR000462 | CDP-alcohol phosphatidyltransferase | 5 | 76 | 9.5E-16 |
| Gene3D | G3DSA:1.20.120.1760 | 5 | 185 | 1.6E-11 | |||
| TIGRFAM | TIGR00560 | pgsA: CDP-diacylglycerol--glycerol-3-phosphate 3-phosphatidyltransferase | IPR004570 | CDP-diacylglycerol--glycerol-3-phosphate 3-phosphatidyltransferase | 4 | 189 | 8.0E-54 |
| ProSitePatterns | PS00379 | CDP-alcohol phosphatidyltransferases signature. | IPR000462 | CDP-alcohol phosphatidyltransferase | 54 | 76 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.