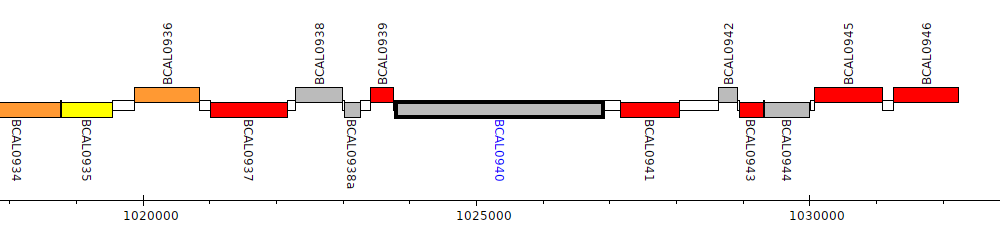
Burkholderia cenocepacia J2315, BCAL0940
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | peptidoglycan biosynthesis IV (Enterococcus faecium) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00550 | Peptidoglycan biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | peptidoglycan biosynthesis V (β-lactam resistance) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | peptidoglycan biosynthesis II (staphylococci) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | peptidoglycan biosynthesis III (mycobacteria) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF56601 | IPR012338 | Beta-lactamase/transpeptidase-like | 469 | 939 | 3.17E-7 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1008 | 1035 | - | ||
| SUPERFAMILY | SSF53955 | IPR023346 | Lysozyme-like domain superfamily | 152 | 370 | 2.02E-9 | |
| SUPERFAMILY | SSF56601 | IPR012338 | Beta-lactamase/transpeptidase-like | 728 | 1004 | 2.48E-23 | |
| Gene3D | G3DSA:3.40.710.10 | 801 | 944 | 1.8E-13 | |||
| Pfam | PF00912 | Transglycosylase | IPR001264 | Glycosyl transferase, family 51 | 150 | 337 | 1.4E-24 |
| Gene3D | G3DSA:1.10.3810.10 | IPR036950 | Penicillin binding protein transglycosylase domain | 136 | 370 | 1.1E-5 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.