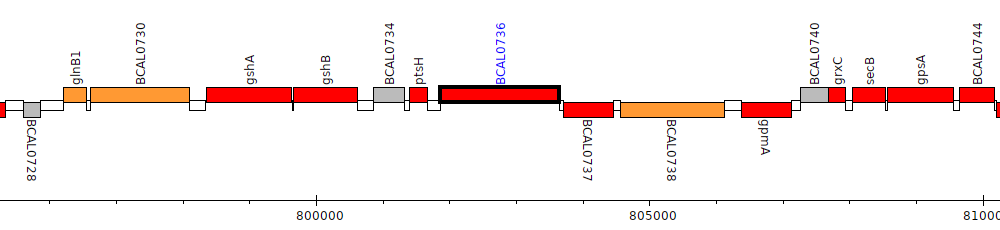
Burkholderia cenocepacia J2315, BCAL0736
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008965 | phosphoenolpyruvate-protein phosphotransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000732
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016310 | phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PS00370
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF51621
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009401 | phosphoenolpyruvate-dependent sugar phosphotransferase system |
Inferred from Sequence Model
Term mapped from: InterPro:PF05524
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016772 | transferase activity, transferring phosphorus-containing groups |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01417
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj02060 | Phosphotransferase system (PTS) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF05524 | PEP-utilising enzyme, N-terminal | IPR008731 | Phosphotransferase system, enzyme I N-terminal | 6 | 129 | 3.7E-29 |
| PIRSF | PIRSF000732 | IPR024692 | Phosphotransferase system, enzyme I | 1 | 584 | 2.8E-210 | |
| SUPERFAMILY | SSF51621 | IPR015813 | Pyruvate/Phosphoenolpyruvate kinase-like domain superfamily | 258 | 557 | 6.81E-100 | |
| Pfam | PF00391 | PEP-utilising enzyme, mobile domain | IPR008279 | PEP-utilising enzyme, mobile domain | 162 | 234 | 7.6E-23 |
| Coils | Coil | 43 | 63 | - | |||
| SUPERFAMILY | SSF47831 | IPR036618 | PtsI, HPr-binding domain superfamily | 28 | 146 | 6.67E-31 | |
| ProSitePatterns | PS00742 | PEP-utilizing enzymes signature 2. | IPR023151 | PEP-utilising enzyme, conserved site | 458 | 476 | - |
| Gene3D | G3DSA:3.50.30.10 | 7 | 242 | 1.2E-74 | |||
| PRINTS | PR01736 | Phosphoenolpyruvate-protein phosphotransferase signature | 511 | 523 | 1.6E-26 | ||
| TIGRFAM | TIGR01417 | PTS_I_fam: phosphoenolpyruvate-protein phosphotransferase | IPR006318 | Phosphotransferase system, enzyme I-like | 5 | 574 | 1.5E-164 |
| Gene3D | G3DSA:3.20.20.60 | IPR040442 | Pyruvate kinase-like domain superfamily | 243 | 583 | 1.0E-111 | |
| Gene3D | G3DSA:1.10.274.10 | IPR036618 | PtsI, HPr-binding domain superfamily | 24 | 149 | 1.2E-74 | |
| PRINTS | PR01736 | Phosphoenolpyruvate-protein phosphotransferase signature | 458 | 473 | 1.6E-26 | ||
| ProSitePatterns | PS00370 | PEP-utilizing enzymes phosphorylation site signature. | IPR018274 | PEP-utilising enzyme, active site | 193 | 204 | - |
| SUPERFAMILY | SSF52009 | IPR036637 | Phosphohistidine domain superfamily | 128 | 254 | 9.68E-31 | |
| PRINTS | PR01736 | Phosphoenolpyruvate-protein phosphotransferase signature | 475 | 490 | 1.6E-26 | ||
| Pfam | PF02896 | PEP-utilising enzyme, TIM barrel domain | IPR000121 | PEP-utilising enzyme, C-terminal | 261 | 551 | 1.9E-108 |
| PRINTS | PR01736 | Phosphoenolpyruvate-protein phosphotransferase signature | 302 | 321 | 1.6E-26 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.