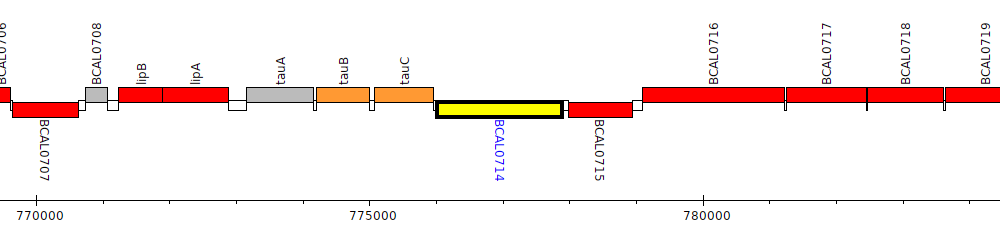
Burkholderia cenocepacia J2315, BCAL0714
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006751 | glutathione catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00066
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0036374 | glutathione hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00066
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00480 | Glutathione metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00430 | Taurine and hypotaurine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | felinine and 3-methyl-3-sulfanylbutan-1-ol biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | glutathione-mediated detoxification I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 4-hydroxy-2-nonenal detoxification | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00460 | Cyanoamino acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | glutathione-mediated detoxification II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00430 | Taurine and hypotaurine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00460 | Cyanoamino acid metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | hypoglycin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | glutathione degradation (DUG pathway - yeast) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00480 | Glutathione metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | γ-glutamyl cycle | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF56235 | IPR029055 | Nucleophile aminohydrolases, N-terminal | 66 | 623 | 4.01E-156 | |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 497 | 512 | 4.2E-65 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 186 | 205 | 4.2E-65 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 523 | 540 | 4.2E-65 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 439 | 457 | 4.2E-65 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 287 | 303 | 4.2E-65 | ||
| Gene3D | G3DSA:3.60.20.40 | 439 | 624 | 2.2E-51 | |||
| TIGRFAM | TIGR00066 | g_glut_trans: gamma-glutamyltransferase | IPR000101 | Gamma-glutamyltranspeptidase | 74 | 616 | 4.0E-112 |
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 461 | 479 | 4.2E-65 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 168 | 186 | 4.2E-65 | ||
| Pfam | PF01019 | Gamma-glutamyltranspeptidase | 88 | 619 | 3.2E-148 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 95 | 120 | 4.2E-65 | ||
| PRINTS | PR01210 | Gamma-glutamyltranspeptidase signature | 317 | 336 | 4.2E-65 | ||
| Gene3D | G3DSA:1.10.246.130 | 303 | 435 | 2.7E-34 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.