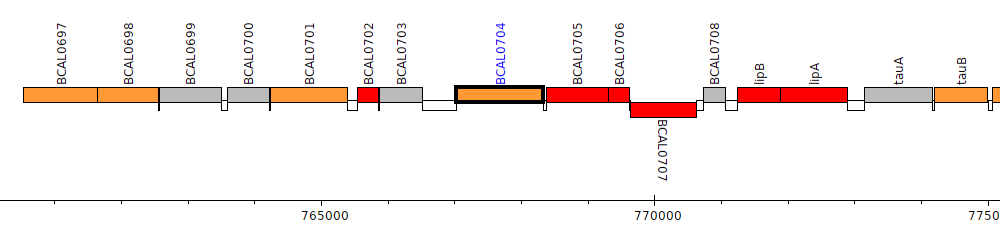
Burkholderia cenocepacia J2315, BCAL0704
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004180 | carboxypeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF69189
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009002 | serine-type D-Ala-D-Ala carboxypeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07943
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:PF07943
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00550 | Peptidoglycan biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00550 | Peptidoglycan biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | peptidoglycan biosynthesis IV (Enterococcus faecium) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | peptidoglycan biosynthesis II (staphylococci) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF69189 | IPR015956 | Penicillin-binding protein, C-terminal domain superfamily | 327 | 419 | 2.55E-24 | |
| PRINTS | PR00725 | D-Ala-D-Ala carboxypeptidase 1 (S11) family signature | IPR018044 | Peptidase S11, D-alanyl-D-alanine carboxypeptidase A | 188 | 201 | 3.2E-18 |
| Gene3D | G3DSA:3.40.710.10 | 47 | 325 | 7.2E-89 | |||
| Pfam | PF07943 | Penicillin-binding protein 5, C-terminal domain | IPR012907 | Peptidase S11, D-Ala-D-Ala carboxypeptidase A, C-terminal | 327 | 417 | 1.9E-20 |
| PRINTS | PR00725 | D-Ala-D-Ala carboxypeptidase 1 (S11) family signature | IPR018044 | Peptidase S11, D-alanyl-D-alanine carboxypeptidase A | 106 | 117 | 3.2E-18 |
| SUPERFAMILY | SSF56601 | IPR012338 | Beta-lactamase/transpeptidase-like | 65 | 326 | 9.45E-77 | |
| Pfam | PF00768 | D-alanyl-D-alanine carboxypeptidase | IPR001967 | Peptidase S11, D-alanyl-D-alanine carboxypeptidase A, N-terminal | 76 | 307 | 1.1E-75 |
| PRINTS | PR00725 | D-Ala-D-Ala carboxypeptidase 1 (S11) family signature | IPR018044 | Peptidase S11, D-alanyl-D-alanine carboxypeptidase A | 161 | 178 | 3.2E-18 |
| Gene3D | G3DSA:2.60.410.10 | IPR037167 | D-Ala-D-Ala carboxypeptidase, C-terminal domain superfamily | 327 | 417 | 3.8E-26 | |
| SMART | SM00936 | IPR012907 | Peptidase S11, D-Ala-D-Ala carboxypeptidase A, C-terminal | 327 | 417 | 7.9E-32 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.