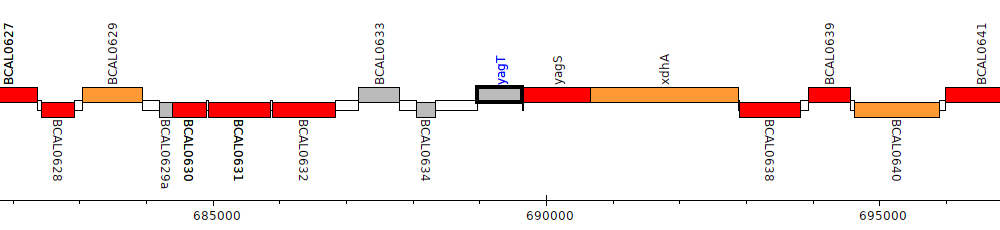
Burkholderia cenocepacia J2315, BCAL0635 (yagT)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF54292
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051537 | 2 iron, 2 sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00197
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF01799
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01799
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF54292
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01799
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF01799 | [2Fe-2S] binding domain | IPR002888 | [2Fe-2S]-binding | 128 | 216 | 1.7E-28 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 29 | 53 | - | ||
| ProSiteProfiles | PS51085 | 2Fe-2S ferredoxin-type iron-sulfur binding domain profile. | IPR001041 | 2Fe-2S ferredoxin-type iron-sulfur binding domain | 56 | 132 | 11.741 |
| Gene3D | G3DSA:3.10.20.30 | IPR012675 | Beta-grasp domain superfamily | 48 | 131 | 1.9E-31 | |
| ProSitePatterns | PS00197 | 2Fe-2S ferredoxin-type iron-sulfur binding region signature. | IPR006058 | 2Fe-2S ferredoxin, iron-sulphur binding site | 94 | 102 | - |
| ProSiteProfiles | PS51318 | Twin arginine translocation (Tat) signal profile. | IPR006311 | Twin-arginine translocation pathway, signal sequence | 1 | 43 | 8.821 |
| SUPERFAMILY | SSF47741 | IPR036884 | [2Fe-2S]-binding domain superfamily | 139 | 222 | 2.62E-26 | |
| CDD | cd00207 | fer2 | IPR001041 | 2Fe-2S ferredoxin-type iron-sulfur binding domain | 58 | 118 | 1.60352E-6 |
| SUPERFAMILY | SSF54292 | IPR036010 | 2Fe-2S ferredoxin-like superfamily | 56 | 142 | 2.62E-26 | |
| Pfam | PF00111 | 2Fe-2S iron-sulfur cluster binding domain | IPR001041 | 2Fe-2S ferredoxin-type iron-sulfur binding domain | 61 | 115 | 2.6E-8 |
| Gene3D | G3DSA:1.10.150.120 | 132 | 223 | 1.2E-30 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.