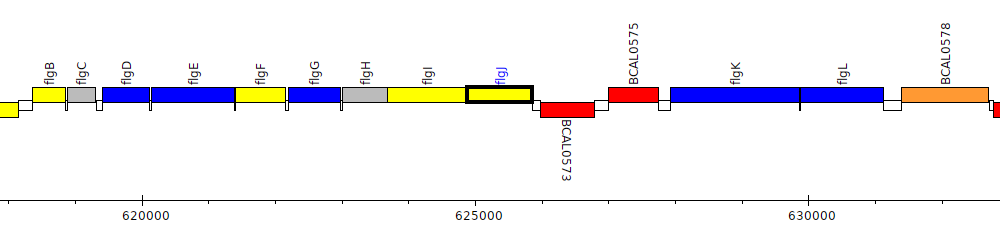
Burkholderia cenocepacia J2315, BCAL0572 (flgJ)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004040 | amidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01832
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0044780 | bacterial-type flagellum assembly |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02541
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016798 | hydrolase activity, acting on glycosyl bonds |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02541
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00047 | IPR002901 | Mannosyl-glycoprotein endo-beta-N-acetylglucosamidase-like domain | 168 | 325 | 7.3E-35 | |
| TIGRFAM | TIGR02541 | flagell_FlgJ: flagellar rod assembly protein/muramidase FlgJ | IPR013377 | Peptidoglycan hydrolase FlgJ | 16 | 324 | 1.8E-100 |
| PRINTS | PR01002 | Flagellar protein FlgJ signature | 78 | 100 | 1.9E-14 | ||
| Gene3D | G3DSA:1.10.530.10 | 181 | 321 | 3.4E-46 | |||
| PRINTS | PR01002 | Flagellar protein FlgJ signature | 57 | 77 | 1.9E-14 | ||
| Pfam | PF10135 | Rod binding protein | IPR019301 | Flagellar protein FlgJ, N-terminal | 54 | 100 | 6.8E-18 |
| PRINTS | PR01002 | Flagellar protein FlgJ signature | 10 | 30 | 1.9E-14 | ||
| PRINTS | PR01002 | Flagellar protein FlgJ signature | 32 | 55 | 1.9E-14 | ||
| Gene3D | G3DSA:2.10.70.40 | 215 | 267 | 3.4E-46 | |||
| Pfam | PF01832 | Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase | IPR002901 | Mannosyl-glycoprotein endo-beta-N-acetylglucosamidase-like domain | 189 | 324 | 3.7E-21 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.