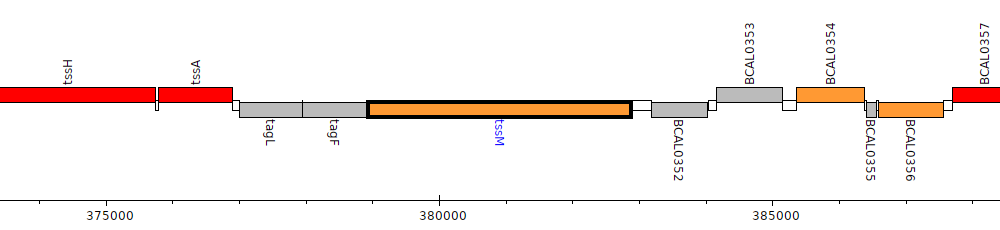
Burkholderia cenocepacia J2315, BCAL0351 (tssM)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0033103 | protein secretion by the type VI secretion system | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
18316384 | Reviewed by curator |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj03070 | Bacterial secretion system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 129 | 278 | 8.99E-7 | |
| TIGRFAM | TIGR03348 | VI_IcmF: type VI secretion protein IcmF | IPR017731 | Type VI secretion system, IcmF | 15 | 570 | 9.9E-138 |
| Pfam | PF06761 | Intracellular multiplication and human macrophage-killing | IPR009612 | IcmF-related | 498 | 877 | 3.5E-95 |
| Pfam | PF06744 | Type VI secretion protein IcmF C-terminal | IPR010623 | Type VI secretion system IcmF, C-terminal | 1017 | 1219 | 1.4E-49 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1254 | 1294 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1254 | 1314 | - | ||
| Pfam | PF14331 | ImcF-related N-terminal domain | IPR025743 | IcmF-related, N-terminal domain | 194 | 447 | 2.0E-82 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.