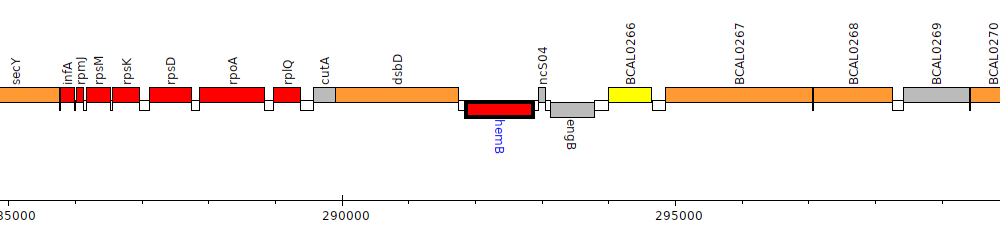
Burkholderia cenocepacia J2315, BCAL0264 (hemB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004655 | porphobilinogen synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00490
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.20.20.70
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00490
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0033014 | tetrapyrrole biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00490
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | tetrapyrrole biosynthesis II (from glycine) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | tetrapyrrole biosynthesis I (from glutamate) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00860 | Porphyrin and chlorophyll metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00860 | Porphyrin and chlorophyll metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF51569 | 3 | 329 | 9.96E-145 | |||
| ProSitePatterns | PS00169 | Delta-aminolevulinic acid dehydratase active site. | IPR030656 | Delta-aminolevulinic acid dehydratase, active site | 247 | 259 | - |
| PRINTS | PR00144 | Delta-aminolevulinic acid dehydratase signature | IPR001731 | Delta-aminolevulinic acid dehydratase | 247 | 263 | 4.7E-57 |
| PRINTS | PR00144 | Delta-aminolevulinic acid dehydratase signature | IPR001731 | Delta-aminolevulinic acid dehydratase | 193 | 212 | 4.7E-57 |
| PRINTS | PR00144 | Delta-aminolevulinic acid dehydratase signature | IPR001731 | Delta-aminolevulinic acid dehydratase | 272 | 287 | 4.7E-57 |
| PRINTS | PR00144 | Delta-aminolevulinic acid dehydratase signature | IPR001731 | Delta-aminolevulinic acid dehydratase | 155 | 174 | 4.7E-57 |
| SMART | SM01004 | IPR001731 | Delta-aminolevulinic acid dehydratase | 4 | 328 | 8.6E-242 | |
| PRINTS | PR00144 | Delta-aminolevulinic acid dehydratase signature | IPR001731 | Delta-aminolevulinic acid dehydratase | 124 | 138 | 4.7E-57 |
| PRINTS | PR00144 | Delta-aminolevulinic acid dehydratase signature | IPR001731 | Delta-aminolevulinic acid dehydratase | 302 | 321 | 4.7E-57 |
| CDD | cd04823 | ALAD_PBGS_aspartate_rich | 7 | 329 | 0.0 | ||
| Pfam | PF00490 | Delta-aminolevulinic acid dehydratase | IPR001731 | Delta-aminolevulinic acid dehydratase | 7 | 326 | 8.7E-140 |
| Gene3D | G3DSA:3.20.20.70 | IPR013785 | Aldolase-type TIM barrel | 1 | 332 | 1.3E-145 | |
| PIRSF | PIRSF001415 | IPR001731 | Delta-aminolevulinic acid dehydratase | 1 | 331 | 1.2E-151 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.