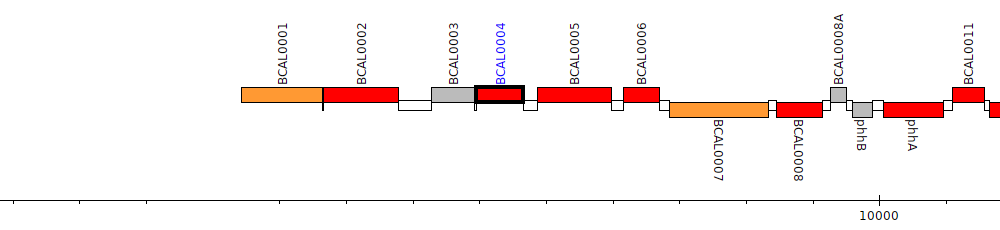
Burkholderia cenocepacia J2315, BCAL0004
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52317 | IPR029062 | Class I glutamine amidotransferase-like | 1 | 232 | 9.32E-45 | |
| CDD | cd01741 | GATase1_1 | 8 | 186 | 3.95314E-60 | ||
| ProSiteProfiles | PS51273 | Glutamine amidotransferase type 1 domain profile. | IPR017926 | Glutamine amidotransferase | 5 | 199 | 16.592 |
| Gene3D | G3DSA:3.40.50.880 | IPR029062 | Class I glutamine amidotransferase-like | 2 | 235 | 1.9E-55 | |
| Pfam | PF00117 | Glutamine amidotransferase class-I | IPR017926 | Glutamine amidotransferase | 20 | 193 | 9.7E-17 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.