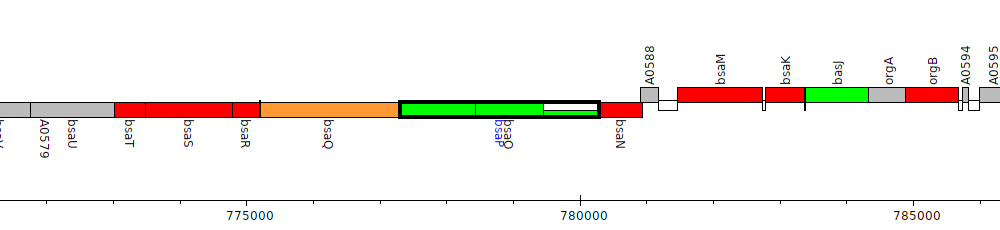
Burkholderia pseudomallei 1710b, BURPS1710b_A0585 (bsaP)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0046903 | secretion |
Inferred from Sequence Model
Term mapped from: InterPro:PF07201
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009986 | cell surface |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02568
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0019867 | outer membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF07201
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0030254 | protein secretion by the type III secretion system |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02568
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009405 | pathogenesis |
Inferred from Sequence Model
Term mapped from: InterPro:PF07201
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009306 | protein secretion |
Inferred from Sequence Model
Term mapped from: InterPro:PR01344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0050709 | negative regulation of protein secretion |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02568
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bpm03070 | Bacterial secretion system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF07201 | HrpJ-like domain | IPR010812 | Hypersensitivity response secretion-like, HrpJ | 674 | 834 | 1.1E-38 |
| Gene3D | G3DSA:3.55.50.30 | 52 | 133 | 5.7E-27 | |||
| PRINTS | PR01344 | Salmonella/Shigella invasion protein E (InvE) signature | IPR003520 | Salmonella/Shigella invasion protein E | 734 | 753 | 2.3E-9 |
| Coils | Coil | 664 | 684 | - | |||
| PRINTS | PR01344 | Salmonella/Shigella invasion protein E (InvE) signature | IPR003520 | Salmonella/Shigella invasion protein E | 798 | 816 | 2.3E-9 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 289 | 354 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 504 | 533 | - | ||
| TIGRFAM | TIGR02568 | LcrE: type III secretion regulator YopN/LcrE/InvE/MxiC | IPR013401 | Type III secretion regulator, YopN/LcrE/InvE/MxiC | 656 | 892 | 7.7E-70 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 396 | 618 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 399 | 473 | - | ||
| Coils | Coil | 391 | 411 | - | |||
| SUPERFAMILY | SSF140591 | 686 | 880 | 4.71E-49 | |||
| PRINTS | PR01344 | Salmonella/Shigella invasion protein E (InvE) signature | IPR003520 | Salmonella/Shigella invasion protein E | 827 | 839 | 2.3E-9 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 489 | 503 | - | ||
| TIGRFAM | TIGR02511 | type_III_tyeA: type III secretion effector delivery regulator, TyeA family | IPR013351 | Type III secretion system effector delivery regulator TyeA-related | 865 | 984 | 4.4E-28 |
| Gene3D | G3DSA:1.10.150.630 | 782 | 871 | 7.9E-34 | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 289 | 318 | - | ||
| Gene3D | G3DSA:3.30.1370.120 | IPR038591 | NolW-like superfamily | 134 | 208 | 1.2E-26 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.