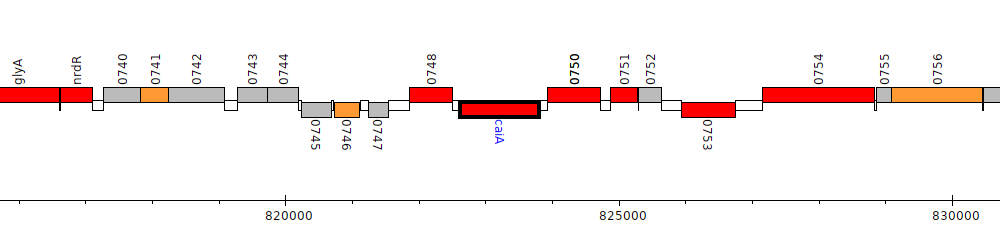
Burkholderia vietnamiensis G4, Bcep1808_0749 (caiA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02771
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02771
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016627 | oxidoreductase activity, acting on the CH-CH group of donors |
Inferred from Sequence Model
Term mapped from: InterPro:PF02771
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003995 | acyl-CoA dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00073
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02771
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02771
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016627 | oxidoreductase activity, acting on the CH-CH group of donors |
Inferred from Sequence Model
Term mapped from: InterPro:PF02771
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003995 | acyl-CoA dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00073
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bvi01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bvi01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bvi00310 | Lysine degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bvi00380 | Tryptophan metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bvi00071 | Fatty acid degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF016578 | 14 | 95 | 0.0027 | |||
| Pfam | PF00441 | Acyl-CoA dehydrogenase, C-terminal domain | IPR009075 | Acyl-CoA dehydrogenase/oxidase C-terminal | 242 | 387 | 6.2E-29 |
| Gene3D | G3DSA:1.10.540.10 | IPR037069 | Acyl-CoA dehydrogenase/oxidase, N-terminal domain superfamily | 1 | 133 | 6.5E-44 | |
| SUPERFAMILY | SSF56645 | IPR009100 | Acyl-CoA dehydrogenase/oxidase, N-terminal and middle domain superfamily | 6 | 241 | 2.88E-79 | |
| SUPERFAMILY | SSF47203 | IPR036250 | Acyl-CoA dehydrogenase-like, C-terminal | 245 | 392 | 2.42E-42 | |
| Pfam | PF02771 | Acyl-CoA dehydrogenase, N-terminal domain | IPR013786 | Acyl-CoA dehydrogenase/oxidase, N-terminal | 19 | 131 | 6.7E-36 |
| CDD | cd01151 | GCD | 6 | 394 | 0.0 | ||
| ProSitePatterns | PS00073 | Acyl-CoA dehydrogenases signature 2. | IPR006089 | Acyl-CoA dehydrogenase, conserved site | 347 | 366 | - |
| Gene3D | G3DSA:2.40.110.10 | 134 | 241 | 1.2E-40 | |||
| Pfam | PF02770 | Acyl-CoA dehydrogenase, middle domain | IPR006091 | Acyl-CoA oxidase/dehydrogenase, central domain | 135 | 229 | 7.3E-21 |
| Gene3D | G3DSA:1.20.140.10 | 243 | 395 | 8.4E-43 | |||
| PIRSF | PIRSF016578 | 148 | 391 | 2.1E-12 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.