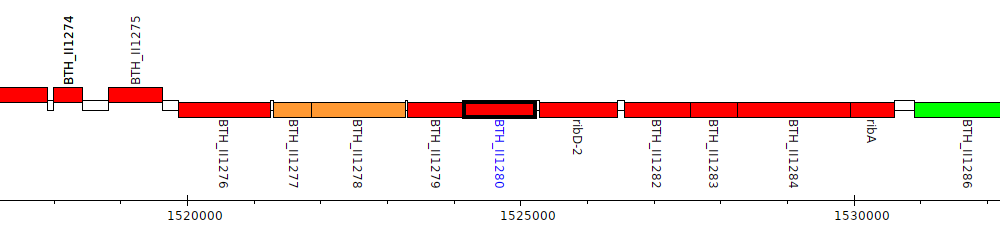
Burkholderia thailandensis E264 ATCC 700388, BTH_II1280
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008168 | methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS51683
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008171 | O-methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00891
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSiteProfiles | PS51683 | SAM-dependent O-methyltransferase class II-type profile. | IPR016461 | O-methyltransferase COMT-type | 26 | 347 | 23.599 |
| Pfam | PF00891 | O-methyltransferase domain | IPR001077 | O-methyltransferase domain | 144 | 312 | 2.3E-16 |
| CDD | cd02440 | AdoMet_MTases | 179 | 282 | 1.65617E-6 | ||
| Gene3D | G3DSA:3.40.50.150 | 103 | 349 | 7.8E-68 | |||
| SUPERFAMILY | SSF53335 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase | 110 | 330 | 2.41E-32 | |
| Pfam | PF16864 | Dimerisation domain | IPR031725 | Acetylserotonin O-methyltransferase, dimerisation domain | 25 | 105 | 8.0E-10 |
| Gene3D | G3DSA:1.10.10.10 | IPR036388 | Winged helix-like DNA-binding domain superfamily | 21 | 102 | 8.2E-36 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.