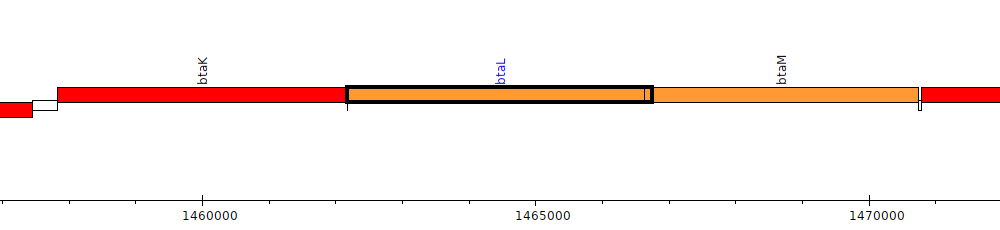
Burkholderia thailandensis E264 ATCC 700388, BTH_II1234 (btaL)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0017000 | antibiotic biosynthetic process | Inferred from Genomic Context | ECO:0000317 genomic context evidence used in manual assertion |
20095633 | Reviewed by curator |
| Molecular Function | GO:0031177 | phosphopantetheine binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00823
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.47.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016740 | transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.366.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.1820 | IPR029058 | Alpha/Beta hydrolase fold | 1395 | 1502 | 8.1E-24 | |
| ProSitePatterns | PS00012 | Phosphopantetheine attachment site. | IPR006162 | Phosphopantetheine attachment site | 1428 | 1443 | - |
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 1119 | 1361 | 2.8E-24 | |
| Pfam | PF16197 | Ketoacyl-synthetase C-terminal extension | IPR032821 | Ketoacyl-synthetase, C-terminal extension | 376 | 461 | 8.2E-9 |
| SMART | SM00827 | Acyl transferase domain in polyketide synthase (PKS) enzymes. | IPR020801 | Polyketide synthase, acyl transferase domain | 511 | 801 | 6.9E-69 |
| Gene3D | G3DSA:3.30.70.3290 | 439 | 847 | 6.1E-95 | |||
| SMART | SM00822 | This enzymatic domain is part of bacterial polyketide synthases and catalyses the first step in the reductive modification of the beta-carbonyl centres in the growing polyketide chain. It uses NADPH to reduce the keto group to a hydroxy group. | 1120 | 1318 | 1.3E-9 | ||
| SMART | SM00825 | Beta-ketoacyl synthase | IPR020841 | Polyketide synthase, beta-ketoacyl synthase domain | 4 | 426 | 3.1E-138 |
| Gene3D | G3DSA:3.40.47.10 | IPR016039 | Thiolase-like | 1 | 438 | 1.7E-143 | |
| ProSitePatterns | PS00606 | Beta-ketoacyl synthases active site. | IPR018201 | Beta-ketoacyl synthase, active site | 159 | 175 | - |
| Pfam | PF02801 | Beta-ketoacyl synthase, C-terminal domain | IPR014031 | Beta-ketoacyl synthase, C-terminal | 255 | 372 | 4.1E-38 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1497 | 1524 | - | ||
| CDD | cd00833 | PKS | 2 | 421 | 5.14425E-165 | ||
| SUPERFAMILY | SSF55048 | IPR016036 | Malonyl-CoA ACP transacylase, ACP-binding | 632 | 692 | 7.06E-8 | |
| Pfam | PF08659 | KR domain | IPR013968 | Polyketide synthase, ketoreductase domain | 1122 | 1316 | 3.7E-23 |
| Pfam | PF00109 | Beta-ketoacyl synthase, N-terminal domain | IPR014030 | Beta-ketoacyl synthase, N-terminal | 1 | 246 | 3.8E-51 |
| Pfam | PF00550 | Phosphopantetheine attachment site | IPR009081 | Phosphopantetheine binding ACP domain | 1406 | 1467 | 6.7E-12 |
| SUPERFAMILY | SSF52151 | IPR016035 | Acyl transferase/acyl hydrolase/lysophospholipase | 509 | 785 | 1.07E-53 | |
| SUPERFAMILY | SSF53901 | IPR016039 | Thiolase-like | 2 | 421 | 4.06E-71 | |
| Gene3D | G3DSA:3.40.50.720 | 849 | 1385 | 7.2E-55 | |||
| SUPERFAMILY | SSF47336 | IPR036736 | ACP-like superfamily | 1397 | 1467 | 1.83E-18 | |
| Pfam | PF00698 | Acyl transferase domain | IPR014043 | Acyl transferase | 511 | 761 | 5.8E-40 |
| SMART | SM00823 | Phosphopantetheine attachment site | IPR020806 | Polyketide synthase, phosphopantetheine-binding domain | 1404 | 1473 | 0.0048 |
| ProSiteProfiles | PS50075 | Carrier protein (CP) domain profile. | IPR009081 | Phosphopantetheine binding ACP domain | 1398 | 1473 | 16.563 |
| Gene3D | G3DSA:3.40.366.10 | IPR001227 | Acyl transferase domain superfamily | 509 | 803 | 6.1E-95 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.