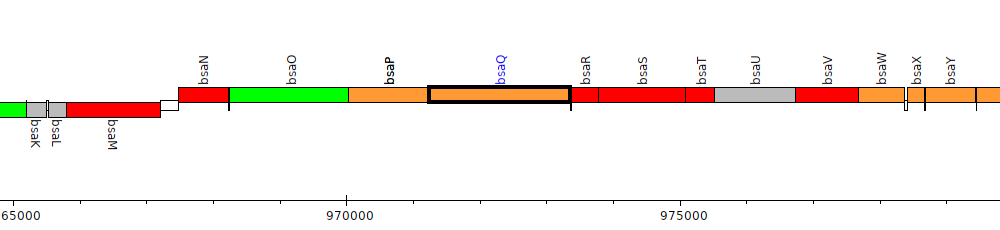
Burkholderia thailandensis E264 ATCC 700388, BTH_II0830 (bsaQ)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PR00949
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01399
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PR00949
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009306 | protein secretion |
Inferred from Sequence Model
Term mapped from: InterPro:PR00949
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015031 | protein transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01399
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009306 | protein secretion |
Inferred from Sequence Model
Term mapped from: InterPro:PR00949
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01399
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015031 | protein transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01399
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bte03070 | Bacterial secretion system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00949 | Type III secretion system inner membrane A protein family signature | IPR001712 | Type III secretion system FHIPEP | 191 | 208 | 1.9E-57 |
| PRINTS | PR00949 | Type III secretion system inner membrane A protein family signature | IPR001712 | Type III secretion system FHIPEP | 80 | 99 | 1.9E-57 |
| Gene3D | G3DSA:1.10.490.80 | IPR042196 | FHIPEP, domain 4 | 604 | 702 | 3.0E-29 | |
| PIRSF | PIRSF005419 | IPR001712 | Type III secretion system FHIPEP | 10 | 702 | 5.9E-212 | |
| PRINTS | PR00949 | Type III secretion system inner membrane A protein family signature | IPR001712 | Type III secretion system FHIPEP | 242 | 256 | 1.9E-57 |
| TIGRFAM | TIGR01399 | hrcV: type III secretion protein, HrcV family | IPR006302 | Type III secretion protein HrcV | 24 | 702 | 2.4E-246 |
| Gene3D | G3DSA:1.10.8.540 | IPR042193 | FHIPEP, domain 3 | 514 | 603 | 2.4E-32 | |
| Coils | Coil | 516 | 536 | - | |||
| Pfam | PF00771 | FHIPEP family | IPR001712 | Type III secretion system FHIPEP | 36 | 694 | 1.1E-214 |
| Gene3D | G3DSA:3.40.5.40 | 445 | 493 | 6.4E-30 | |||
| Coils | Coil | 702 | 702 | - | |||
| ProSitePatterns | PS00994 | Bacterial export FHIPEP family signature. | IPR025505 | FHIPEP conserved site | 147 | 170 | - |
| Gene3D | G3DSA:3.40.30.60 | IPR042194 | FHIPEP, domain 1 | 373 | 512 | 6.4E-30 | |
| PRINTS | PR00949 | Type III secretion system inner membrane A protein family signature | IPR001712 | Type III secretion system FHIPEP | 161 | 179 | 1.9E-57 |
| PRINTS | PR00949 | Type III secretion system inner membrane A protein family signature | IPR001712 | Type III secretion system FHIPEP | 43 | 56 | 1.9E-57 |
| PRINTS | PR00949 | Type III secretion system inner membrane A protein family signature | IPR001712 | Type III secretion system FHIPEP | 142 | 158 | 1.9E-57 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.