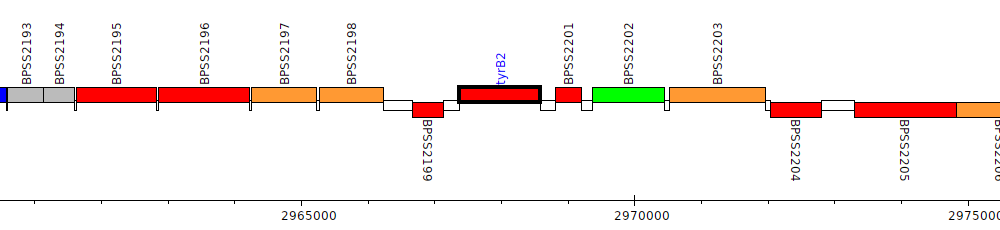
Burkholderia pseudomallei K96243, BPSS2200 (tyrB2)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.640.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PR00799
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030170 | pyridoxal phosphate binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00155
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00155
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008483 | transaminase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00799
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00270 | Cysteine and methionine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | (<i>R</i>)-cysteate degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00350 | Tyrosine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | C4 photosynthetic carbon assimilation cycle, PEPCK type | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00270 | Cysteine and methionine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00960 | Tropane, piperidine and pyridine alkaloid biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00401 | Novobiocin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | sulfolactate degradation III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00350 | Tyrosine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | anaerobic energy metabolism (invertebrates, cytosol) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00950 | Isoquinoline alkaloid biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00220 | Arginine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00360 | Phenylalanine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00401 | Novobiocin biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00250 | Alanine, aspartate and glutamate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00710 | Carbon fixation in photosynthetic organisms | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00330 | Arginine and proline metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | partial TCA cycle (obligate autotrophs) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00360 | Phenylalanine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | coenzyme M biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | C4 photosynthetic carbon assimilation cycle, NAD-ME type | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00799 | Aspartate aminotransferase signature | IPR000796 | Aspartate/other aminotransferase | 206 | 218 | 2.8E-34 |
| SUPERFAMILY | SSF53383 | IPR015424 | Pyridoxal phosphate-dependent transferase | 1 | 397 | 2.62E-118 | |
| Gene3D | G3DSA:3.90.1150.10 | IPR015422 | Pyridoxal phosphate-dependent transferase domain 1 | 5 | 392 | 3.3E-157 | |
| PRINTS | PR00799 | Aspartate aminotransferase signature | IPR000796 | Aspartate/other aminotransferase | 273 | 298 | 2.8E-34 |
| PRINTS | PR00799 | Aspartate aminotransferase signature | IPR000796 | Aspartate/other aminotransferase | 175 | 194 | 2.8E-34 |
| CDD | cd00609 | AAT_like | 29 | 379 | 1.41065E-45 | ||
| Gene3D | G3DSA:3.40.640.10 | IPR015421 | Pyridoxal phosphate-dependent transferase, major domain | 60 | 300 | 3.3E-157 | |
| Pfam | PF00155 | Aminotransferase class I and II | IPR004839 | Aminotransferase, class I/classII | 27 | 392 | 1.6E-81 |
| PRINTS | PR00799 | Aspartate aminotransferase signature | IPR000796 | Aspartate/other aminotransferase | 341 | 359 | 2.8E-34 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.