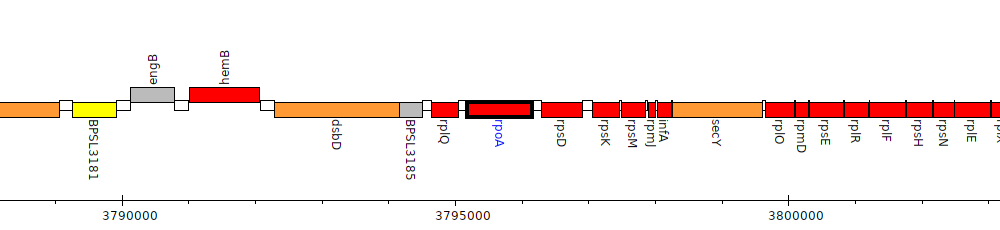
Burkholderia pseudomallei K96243, BPSL3187 (rpoA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006351 | transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PF01193
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003899 | DNA-directed 5'-3' RNA polymerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01193
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00059
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046983 | protein dimerization activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01000
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps00240 | Pyrimidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps03020 | RNA polymerase | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00662 | IPR011263 | DNA-directed RNA polymerase, RpoA/D/Rpb3-type | 20 | 230 | 1.3E-93 | |
| Pfam | PF01000 | RNA polymerase Rpb3/RpoA insert domain | IPR011262 | DNA-directed RNA polymerase, insert domain | 61 | 150 | 1.7E-23 |
| TIGRFAM | TIGR02027 | rpoA: DNA-directed RNA polymerase, alpha subunit | IPR011773 | DNA-directed RNA polymerase, alpha subunit | 21 | 313 | 4.8E-106 |
| Pfam | PF03118 | Bacterial RNA polymerase, alpha chain C terminal domain | IPR011260 | RNA polymerase, alpha subunit, C-terminal | 248 | 305 | 9.3E-25 |
| SUPERFAMILY | SSF55257 | IPR036603 | RNA polymerase, RBP11-like subunit | 5 | 228 | 1.77E-31 | |
| SUPERFAMILY | SSF56553 | IPR036643 | DNA-directed RNA polymerase, insert domain superfamily | 50 | 174 | 5.23E-41 | |
| Pfam | PF01193 | RNA polymerase Rpb3/Rpb11 dimerisation domain | IPR011263 | DNA-directed RNA polymerase, RpoA/D/Rpb3-type | 26 | 224 | 5.4E-22 |
| Gene3D | G3DSA:1.10.150.20 | 242 | 325 | 1.6E-37 | |||
| CDD | cd06928 | RNAP_alpha_NTD | 13 | 227 | 8.62392E-107 | ||
| Gene3D | G3DSA:3.30.1360.10 | IPR036603 | RNA polymerase, RBP11-like subunit | 14 | 223 | 5.3E-81 | |
| Hamap | MF_00059 | DNA-directed RNA polymerase subunit alpha [rpoA]. | IPR011773 | DNA-directed RNA polymerase, alpha subunit | 2 | 314 | 35.125 |
| SUPERFAMILY | SSF47789 | 248 | 320 | 5.23E-27 | |||
| Gene3D | G3DSA:2.170.120.12 | IPR036643 | DNA-directed RNA polymerase, insert domain superfamily | 50 | 173 | 5.3E-81 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.