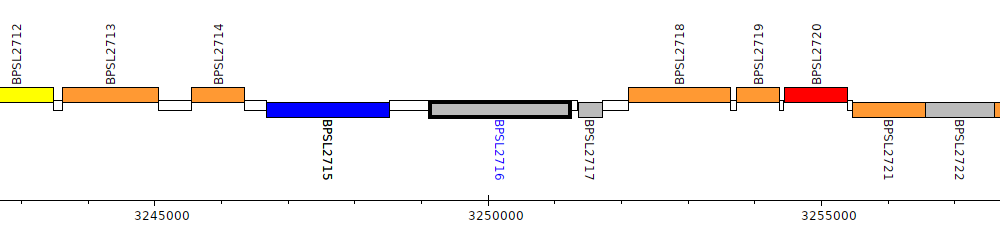
Burkholderia pseudomallei K96243, BPSL2716
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0019605 | butyrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF011409
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005615 | extracellular space |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF011409
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0047989 | hydroxybutyrate-dimer hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF011409
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00650 | Butanoate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bps00650 | Butanoate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF10605 | 3HB-oligomer hydrolase (3HBOH) | IPR016582 | D-(-)-3-hydroxybutyrate oligomer hydrolase, putative | 24 | 698 | 0.0 |
| SUPERFAMILY | SSF53474 | IPR029058 | Alpha/Beta hydrolase fold | 149 | 615 | 4.44E-5 | |
| Hamap | MF_01906 | D-(-)-3-hydroxybutyrate oligomer hydrolase. | IPR016582 | D-(-)-3-hydroxybutyrate oligomer hydrolase, putative | 3 | 699 | 31.861 |
| PIRSF | PIRSF011409 | IPR016582 | D-(-)-3-hydroxybutyrate oligomer hydrolase, putative | 1 | 699 | 0.0 | |
| ProSiteProfiles | PS51257 | Prokaryotic membrane lipoprotein lipid attachment site profile. | 1 | 27 | 5.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.