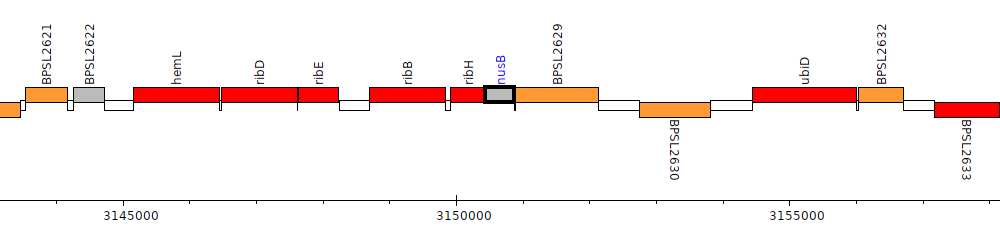
Burkholderia pseudomallei K96243, BPSL2628 (nusB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006353 | DNA-templated transcription, termination |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01951
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PF01029
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Hamap | MF_00073 | Transcription antitermination protein NusB [nusB]. | IPR011605 | NusB antitermination factor | 4 | 135 | 21.31 |
| Gene3D | G3DSA:1.10.940.10 | IPR035926 | NusB-like superfamily | 1 | 139 | 4.2E-42 | |
| TIGRFAM | TIGR01951 | nusB: transcription antitermination factor NusB | IPR011605 | NusB antitermination factor | 6 | 133 | 3.4E-38 |
| SUPERFAMILY | SSF48013 | IPR035926 | NusB-like superfamily | 3 | 135 | 2.62E-40 | |
| Pfam | PF01029 | NusB family | IPR006027 | NusB/RsmB/TIM44 | 8 | 131 | 3.8E-32 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.