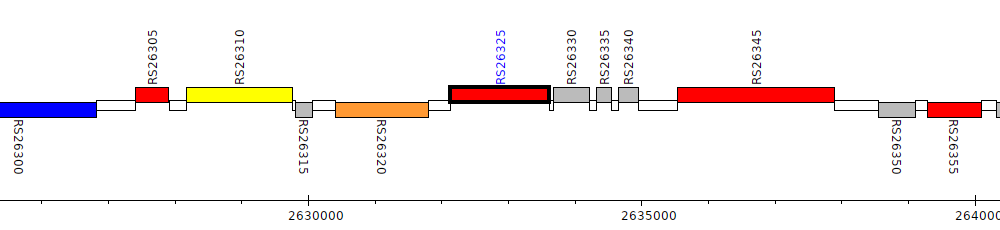
Burkholderia cenocepacia AU 1054, BCEN_RS26325
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PF00158
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0043565 | sequence-specific DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PR01590
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008134 | transcription factor binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00158
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF46689
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00158
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSitePatterns | PS00688 | Sigma-54 interaction domain C-terminal part signature. | IPR025944 | Sigma-54 interaction domain, conserved site | 376 | 385 | - |
| Gene3D | G3DSA:1.10.10.60 | 444 | 495 | 2.5E-11 | |||
| Gene3D | G3DSA:1.10.8.60 | 333 | 409 | 5.4E-21 | |||
| Pfam | PF08448 | PAS fold | IPR013656 | PAS fold-4 | 27 | 119 | 3.8E-8 |
| SUPERFAMILY | SSF55785 | IPR035965 | PAS domain superfamily | 15 | 118 | 5.14E-11 | |
| CDD | cd00009 | AAA | 170 | 324 | 2.16645E-22 | ||
| PRINTS | PR01590 | FIS bacterial regulatory protein HTH signature | IPR002197 | DNA binding HTH domain, Fis-type | 457 | 474 | 5.3E-5 |
| SMART | SM00382 | IPR003593 | AAA+ ATPase domain | 181 | 326 | 2.0E-9 | |
| Gene3D | G3DSA:3.40.50.300 | 147 | 332 | 4.7E-61 | |||
| ProSiteProfiles | PS50045 | Sigma-54 interaction domain profile. | IPR002078 | RNA polymerase sigma factor 54 interaction domain | 161 | 392 | 62.421 |
| SUPERFAMILY | SSF46689 | IPR009057 | Homeobox-like domain superfamily | 444 | 494 | 6.09E-11 | |
| Coils | Coil | 134 | 154 | - | |||
| Gene3D | G3DSA:3.30.450.20 | 13 | 119 | 3.7E-9 | |||
| ProSitePatterns | PS00675 | Sigma-54 interaction domain ATP-binding region A signature. | IPR025662 | Sigma-54 interaction domain, ATP-binding site 1 | 185 | 198 | - |
| PRINTS | PR01590 | FIS bacterial regulatory protein HTH signature | IPR002197 | DNA binding HTH domain, Fis-type | 474 | 494 | 5.3E-5 |
| Pfam | PF00158 | Sigma-54 interaction domain | IPR002078 | RNA polymerase sigma factor 54 interaction domain | 162 | 329 | 2.2E-62 |
| Pfam | PF02954 | Bacterial regulatory protein, Fis family | IPR002197 | DNA binding HTH domain, Fis-type | 451 | 490 | 3.0E-8 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 161 | 405 | 1.49E-64 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.