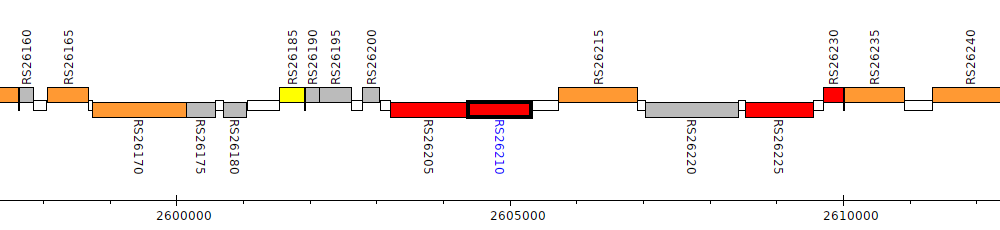
Burkholderia cenocepacia AU 1054, BCEN_RS26210
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02401
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0050992 | dimethylallyl diphosphate biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02401
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051745 | 4-hydroxy-3-methylbut-2-en-1-yl diphosphate reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02401
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019288 | isopentenyl diphosphate biosynthetic process, methylerythritol 4-phosphate pathway |
Inferred from Sequence Model
Term mapped from: InterPro:PF02401
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.1010.20 | 6 | 278 | 6.9E-108 | |||
| Hamap | MF_00191 | 4-hydroxy-3-methylbut-2-enyl diphosphate reductase [ispH]. | IPR003451 | 4-hydroxy-3-methylbut-2-enyl diphosphate reductase | 1 | 283 | 37.31 |
| Pfam | PF02401 | LytB protein | IPR003451 | 4-hydroxy-3-methylbut-2-enyl diphosphate reductase | 3 | 279 | 4.0E-90 |
| Gene3D | G3DSA:3.40.50.11270 | 12 | 96 | 6.9E-108 | |||
| CDD | cd13944 | lytB_ispH | IPR003451 | 4-hydroxy-3-methylbut-2-enyl diphosphate reductase | 3 | 279 | 1.41819E-142 |
| TIGRFAM | TIGR00216 | ispH_lytB: 4-hydroxy-3-methylbut-2-enyl diphosphate reductase | IPR003451 | 4-hydroxy-3-methylbut-2-enyl diphosphate reductase | 3 | 281 | 1.8E-97 |
| Gene3D | G3DSA:3.40.1010.20 | 97 | 193 | 6.9E-108 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.