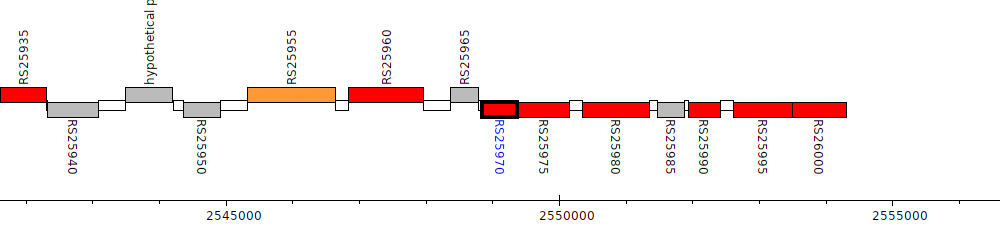
Burkholderia cenocepacia AU 1054, BCEN_RS25970
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006979 | response to oxidative stress |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03561
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:2.20.25.10 | 18 | 67 | 1.0E-7 | |||
| Pfam | PF02566 | OsmC-like protein | IPR003718 | OsmC/Ohr family | 63 | 157 | 4.9E-15 |
| Gene3D | G3DSA:3.30.300.20 | IPR015946 | K homology domain-like, alpha/beta | 69 | 162 | 1.2E-20 | |
| SUPERFAMILY | SSF82784 | IPR036102 | OsmC/Ohr superfamily | 18 | 158 | 1.57E-31 | |
| TIGRFAM | TIGR03561 | organ_hyd_perox: peroxiredoxin, Ohr subfamily | IPR019953 | Organic hydroperoxide resistance protein famiy | 24 | 156 | 3.3E-32 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.