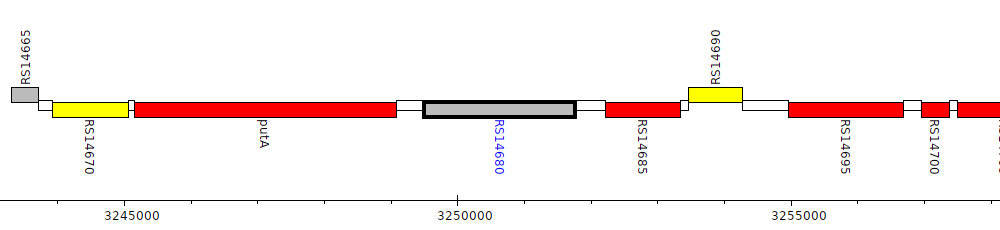
Burkholderia cenocepacia AU 1054, BCEN_RS14680
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00595
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0032508 | DNA duplex unwinding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00595
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00270
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006260 | DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00595
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00270
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003678 | DNA helicase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00595
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.300 | 201 | 397 | 4.8E-67 | |||
| Pfam | PF00270 | DEAD/DEAH box helicase | IPR011545 | DEAD/DEAH box helicase domain | 212 | 378 | 4.2E-12 |
| SUPERFAMILY | SSF52540 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 227 | 623 | 3.84E-38 | |
| TIGRFAM | TIGR00595 | priA: primosomal protein N' | IPR005259 | Primosomal protein N' | 232 | 751 | 3.1E-166 |
| Hamap | MF_00983 | Probable primosomal protein N' [priA]. | IPR005259 | Primosomal protein N' | 8 | 753 | 22.601 |
| SMART | SM00487 | IPR014001 | Helicase superfamily 1/2, ATP-binding domain | 207 | 411 | 9.3E-14 | |
| Pfam | PF18074 | Primosomal protein N C-terminal domain | IPR041236 | Primosomal protein N, C-terminal domain | 656 | 750 | 8.5E-18 |
| CDD | cd18804 | SF2_C_priA | 408 | 650 | 7.77931E-112 | ||
| ProSiteProfiles | PS51194 | Superfamilies 1 and 2 helicase C-terminal domain profile. | IPR001650 | Helicase, C-terminal | 495 | 647 | 8.363 |
| SMART | SM00490 | IPR001650 | Helicase, C-terminal | 516 | 613 | 4.2E-10 | |
| CDD | cd17929 | DEXHc_priA | 215 | 398 | 1.36698E-76 | ||
| Pfam | PF17764 | 3??? DNA-binding domain (3???BD) | IPR041222 | Primosomal protein N', 3' DNA-binding domain | 7 | 102 | 1.3E-20 |
| Gene3D | G3DSA:1.10.8.630 | IPR042115 | Primosomal protein N', 3' DNA-binding domain superfamily | 5 | 111 | 7.8E-24 | |
| Coils | Coil | 432 | 452 | - | |||
| Pfam | PF18319 | PriA DNA helicase Cys-rich region (CRR) domain | IPR040498 | PriA DNA helicase, Cys-rich region (CRR) domain | 469 | 494 | 6.3E-9 |
| Pfam | PF00271 | Helicase conserved C-terminal domain | IPR001650 | Helicase, C-terminal | 528 | 613 | 3.1E-7 |
| ProSiteProfiles | PS51192 | Superfamilies 1 and 2 helicase ATP-binding type-1 domain profile. | IPR014001 | Helicase superfamily 1/2, ATP-binding domain | 222 | 395 | 16.193 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.