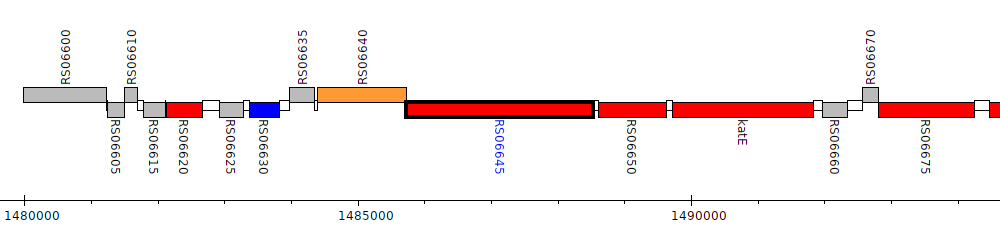
Burkholderia cenocepacia AU 1054, BCEN_RS06645
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006269 | DNA replication, synthesis of RNA primer |
Inferred from Sequence Model
Term mapped from: InterPro:PF01896
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003896 | DNA primase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01896
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006281 | DNA repair |
Inferred from Sequence Model
Term mapped from: InterPro:PS50160
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006310 | DNA recombination |
Inferred from Sequence Model
Term mapped from: InterPro:PS50160
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003910 | DNA ligase (ATP) activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50160
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50160
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF01068 | ATP dependent DNA ligase domain | IPR012310 | DNA ligase, ATP-dependent, central | 282 | 455 | 8.7E-21 |
| TIGRFAM | TIGR02779 | NHEJ_ligase_lig: DNA ligase D, ligase domain | IPR014146 | DNA ligase D, ligase domain | 270 | 584 | 2.7E-97 |
| ProSiteProfiles | PS50160 | ATP-dependent DNA ligase family profile. | IPR012310 | DNA ligase, ATP-dependent, central | 363 | 455 | 12.244 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 187 | 258 | - | ||
| TIGRFAM | TIGR02777 | LigD_PE_dom: DNA ligase D, 3'-phosphoesterase domain | IPR014144 | DNA ligase D, 3'-phosphoesterase domain | 29 | 183 | 5.2E-63 |
| Pfam | PF01896 | DNA primase small subunit | IPR002755 | DNA primase, small subunit | 764 | 885 | 2.3E-6 |
| Gene3D | G3DSA:2.40.50.140 | 457 | 586 | 1.4E-36 | |||
| Gene3D | G3DSA:3.30.470.30 | 293 | 412 | 1.5E-41 | |||
| Pfam | PF04679 | ATP dependent DNA ligase C terminal region | IPR012309 | DNA ligase, ATP-dependent, C-terminal | 474 | 578 | 3.0E-23 |
| CDD | cd07971 | OBF_DNA_ligase_LigD | 458 | 583 | 6.51226E-48 | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 216 | 250 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 48 | - | ||
| TIGRFAM | TIGR02776 | NHEJ_ligase_prk: DNA ligase D | IPR014143 | DNA ligase D | 279 | 919 | 2.4E-207 |
| Pfam | PF13298 | DNA polymerase Ligase (LigD) | IPR014144 | DNA ligase D, 3'-phosphoesterase domain | 57 | 164 | 2.5E-39 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 35 | - | ||
| Gene3D | G3DSA:3.90.920.30 | 653 | 935 | 3.0E-106 | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 582 | 654 | - | ||
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 458 | 585 | 2.83E-26 | |
| CDD | cd04862 | PaeLigD_Pol_like | IPR033651 | LigD polymerase domain, PaeLigD-type | 663 | 888 | 5.48779E-128 |
| SUPERFAMILY | SSF56091 | 252 | 456 | 2.59E-44 | |||
| Gene3D | G3DSA:3.30.1490.70 | 280 | 453 | 1.5E-41 | |||
| TIGRFAM | TIGR02778 | ligD_pol: DNA ligase D, polymerase domain | IPR014145 | DNA ligase D, polymerase domain | 646 | 888 | 1.9E-90 |
| CDD | cd07906 | Adenylation_DNA_ligase_LigD_LigC | 268 | 457 | 8.63487E-77 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.