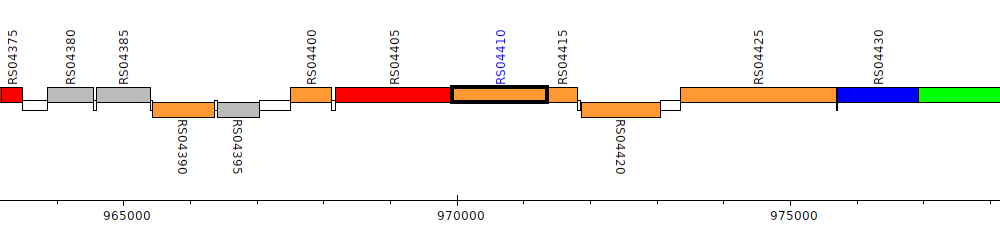
Burkholderia cenocepacia AU 1054, BCEN_RS04410
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006470 | protein dephosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:SM00195
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008138 | protein tyrosine/serine/threonine phosphatase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00195
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016791 | phosphatase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50056
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016311 | dephosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PS50056
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00782 | Dual specificity phosphatase, catalytic domain | IPR000340 | Dual specificity phosphatase, catalytic domain | 347 | 461 | 1.8E-12 |
| CDD | cd14527 | DSP_bac | 324 | 461 | 3.55211E-30 | ||
| SMART | SM00195 | IPR020422 | Dual specificity protein phosphatase domain | 327 | 464 | 8.9E-7 | |
| CDD | cd03386 | PAP2_Aur1_like | 46 | 220 | 1.7997E-19 | ||
| SUPERFAMILY | SSF52799 | IPR029021 | Protein-tyrosine phosphatase-like | 309 | 465 | 2.12E-22 | |
| Gene3D | G3DSA:3.90.190.10 | IPR029021 | Protein-tyrosine phosphatase-like | 329 | 466 | 5.7E-19 | |
| ProSiteProfiles | PS50056 | Tyrosine specific protein phosphatases family profile. | IPR000387 | Tyrosine specific protein phosphatases domain | 387 | 456 | 11.153 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.