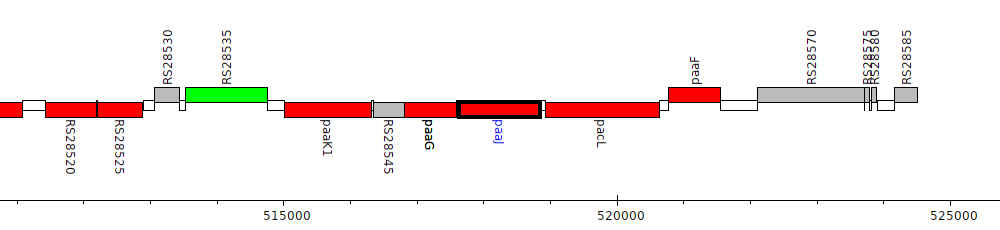
Burkholderia cenocepacia K56-2Valvano, BURCENK562V_RS28555 (paaJ)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0033812 | 3-oxoadipyl-CoA thiolase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P0C7L2
|
ECO:00000060 |
17259607 | Reviewed by curator |
| Biological Process | GO:0010124 | phenylacetate catabolic process |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P0C7L2
|
ECO:00000060 |
12846838 | Reviewed by curator |
| Molecular Function | GO:0016740 | transferase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P0C7L2
|
ECO:00000060 |
17259607 | Reviewed by curator |
| Molecular Function | GO:0016747 | transferase activity, transferring acyl groups other than amino-acyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:PS00098
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.47.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019619 | 3,4-dihydroxybenzoate catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02430
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.47.10 | IPR016039 | Thiolase-like | 6 | 281 | 1.2E-152 | |
| SUPERFAMILY | SSF53901 | IPR016039 | Thiolase-like | 3 | 275 | 9.27E-79 | |
| TIGRFAM | TIGR01930 | AcCoA-C-Actrans: acetyl-CoA C-acyltransferase | IPR002155 | Thiolase | 6 | 398 | 5.0E-133 |
| Gene3D | G3DSA:3.40.47.10 | IPR016039 | Thiolase-like | 141 | 397 | 1.2E-152 | |
| PIRSF | PIRSF000429 | IPR002155 | Thiolase | 1 | 400 | 3.7E-132 | |
| CDD | cd00751 | thiolase | IPR002155 | Thiolase | 5 | 399 | 0.0 |
| SUPERFAMILY | SSF53901 | IPR016039 | Thiolase-like | 277 | 399 | 1.11E-44 | |
| ProSitePatterns | PS00098 | Thiolases acyl-enzyme intermediate signature. | IPR020615 | Thiolase, acyl-enzyme intermediate active site | 86 | 104 | - |
| Pfam | PF00108 | Thiolase, N-terminal domain | IPR020616 | Thiolase, N-terminal | 5 | 267 | 1.3E-72 |
| ProSitePatterns | PS00737 | Thiolases signature 2. | IPR020613 | Thiolase, conserved site | 346 | 362 | - |
| TIGRFAM | TIGR02430 | pcaF: 3-oxoadipyl-CoA thiolase | IPR012793 | Beta-ketoadipyl CoA thiolase | 3 | 400 | 4.3E-222 |
| Pfam | PF02803 | Thiolase, C-terminal domain | IPR020617 | Thiolase, C-terminal | 277 | 399 | 2.2E-43 |
| ProSitePatterns | PS00099 | Thiolases active site. | IPR020610 | Thiolase, active site | 381 | 394 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.