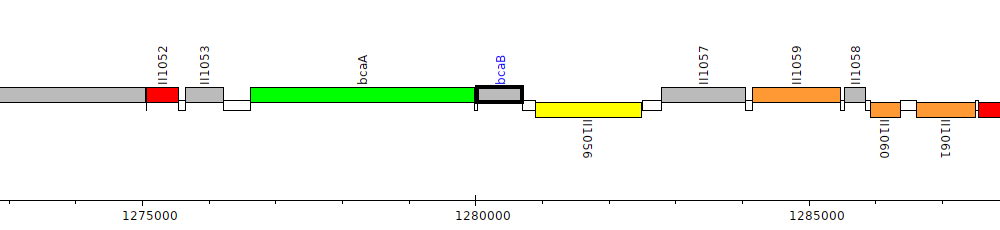
Burkholderia pseudomallei 1026b, BP1026B_II1055 (bcaB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0044407 | single-species biofilm formation in or on host organism | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
23340315 | Reviewed by curator |
| Biological Process | GO:0035635 | entry of bacterium into host cell | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
23340315 | Reviewed by curator |
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF13640
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF13640
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF13640
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF13640
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF13640 | 2OG-Fe(II) oxygenase superfamily | IPR005123 | Oxoglutarate/iron-dependent dioxygenase | 142 | 216 | 3.6E-6 |
| Gene3D | G3DSA:2.60.120.620 | 32 | 222 | 4.9E-17 | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 32 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 8 | 22 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.