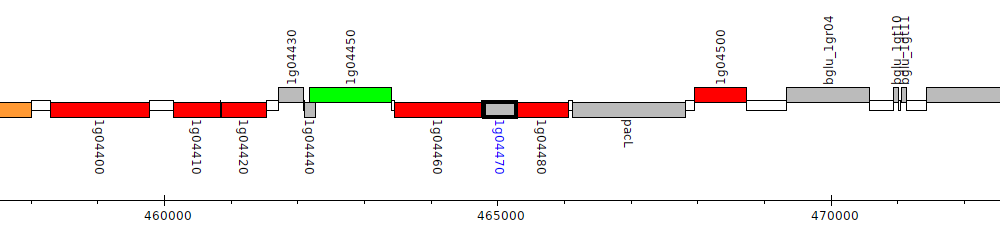
Burkholderia glumae BGR1, bglu_1g04470
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016790 | thiolester hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02286
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bgl00360 | Phenylalanine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR02286 | PaaD: phenylacetic acid degradation protein PaaD | IPR011973 | Phenylacetic acid degradation protein PaaD | 38 | 150 | 1.2E-47 |
| SUPERFAMILY | SSF54637 | IPR029069 | HotDog domain superfamily | 19 | 145 | 2.45E-30 | |
| Pfam | PF03061 | Thioesterase superfamily | IPR006683 | Thioesterase domain | 67 | 140 | 6.8E-12 |
| CDD | cd03443 | PaaI_thioesterase | 39 | 149 | 1.10135E-26 | ||
| TIGRFAM | TIGR00369 | unchar_dom_1: uncharacterized domain 1 | IPR003736 | Phenylacetic acid degradation-related domain | 36 | 148 | 3.7E-25 |
| Gene3D | G3DSA:3.10.129.10 | 21 | 159 | 1.8E-50 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.