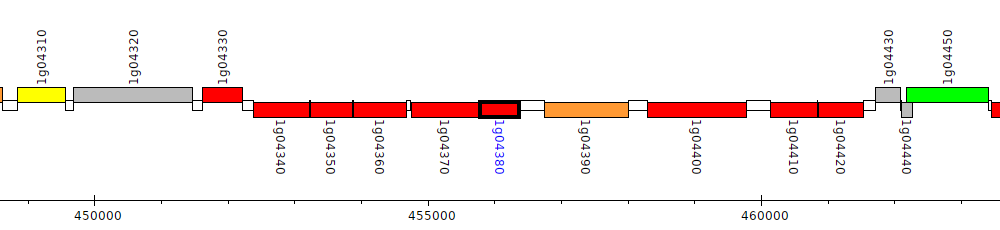
Burkholderia glumae BGR1, bglu_1g04380
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | 4-hydroxy-2(1<i>H</i>)-quinolone biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bgl01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bgl00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00405 | Phenazine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bgl01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | acridone alkaloid biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bgl01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSiteProfiles | PS51273 | Glutamine amidotransferase type 1 domain profile. | IPR017926 | Glutamine amidotransferase | 1 | 194 | 29.362 |
| PRINTS | PR00096 | Glutamine amidotransferase superfamily signature | 164 | 177 | 5.3E-14 | ||
| PRINTS | PR00097 | Anthranilate synthase component II signature | 119 | 131 | 6.0E-35 | ||
| SUPERFAMILY | SSF52317 | IPR029062 | Class I glutamine amidotransferase-like | 1 | 189 | 3.8E-57 | |
| PRINTS | PR00099 | Carbamoyl-phosphate synthase protein GATase domain signature | 44 | 58 | 1.8E-6 | ||
| PRINTS | PR00097 | Anthranilate synthase component II signature | 2 | 16 | 6.0E-35 | ||
| PRINTS | PR00097 | Anthranilate synthase component II signature | 74 | 85 | 6.0E-35 | ||
| PRINTS | PR00097 | Anthranilate synthase component II signature | 47 | 56 | 6.0E-35 | ||
| PRINTS | PR00096 | Glutamine amidotransferase superfamily signature | 74 | 85 | 5.3E-14 | ||
| PRINTS | PR00097 | Anthranilate synthase component II signature | 99 | 107 | 6.0E-35 | ||
| PRINTS | PR00099 | Carbamoyl-phosphate synthase protein GATase domain signature | 74 | 90 | 1.8E-6 | ||
| Pfam | PF00117 | Glutamine amidotransferase class-I | IPR017926 | Glutamine amidotransferase | 4 | 187 | 1.9E-55 |
| CDD | cd01743 | GATase1_Anthranilate_Synthase | 2 | 185 | 9.41911E-117 | ||
| Gene3D | G3DSA:3.40.50.880 | IPR029062 | Class I glutamine amidotransferase-like | 1 | 190 | 2.2E-65 | |
| PRINTS | PR00096 | Glutamine amidotransferase superfamily signature | 47 | 56 | 5.3E-14 | ||
| TIGRFAM | TIGR00566 | trpG_papA: glutamine amidotransferase of anthranilate synthase or aminodeoxychorismate synthase | IPR006221 | Anthranilate synthase/para-aminobenzoate synthase like domain | 1 | 186 | 4.2E-79 |
| PRINTS | PR00097 | Anthranilate synthase component II signature | 164 | 177 | 6.0E-35 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.