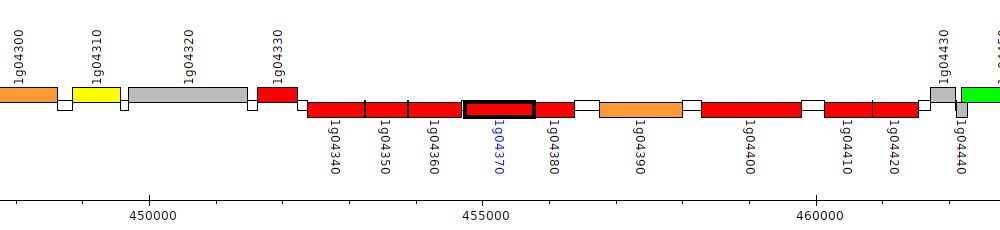
Burkholderia glumae BGR1, bglu_1g04370
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004048 | anthranilate phosphoribosyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00211
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000162 | tryptophan biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00211
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016757 | transferase activity, transferring glycosyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:PF00591
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bgl01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bgl00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bgl01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bgl01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00400 | Phenylalanine, tyrosine and tryptophan biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR01245 | trpD: anthranilate phosphoribosyltransferase | IPR005940 | Anthranilate phosphoribosyl transferase | 8 | 335 | 5.4E-126 |
| SUPERFAMILY | SSF52418 | IPR035902 | Nucleoside phosphorylase/phosphoribosyltransferase catalytic domain superfamily | 77 | 337 | 5.76E-97 | |
| Gene3D | G3DSA:3.40.1030.10 | IPR035902 | Nucleoside phosphorylase/phosphoribosyltransferase catalytic domain superfamily | 72 | 340 | 4.8E-109 | |
| Hamap | MF_00211 | Anthranilate phosphoribosyltransferase [trpD]. | IPR005940 | Anthranilate phosphoribosyl transferase | 6 | 336 | 37.133 |
| Pfam | PF00591 | Glycosyl transferase family, a/b domain | IPR000312 | Glycosyl transferase, family 3 | 76 | 326 | 1.3E-93 |
| Gene3D | G3DSA:1.20.970.10 | 1 | 71 | 7.7E-31 | |||
| SUPERFAMILY | SSF47648 | IPR036320 | Glycosyl transferase family 3, N-terminal domain superfamily | 3 | 70 | 3.4E-16 | |
| Pfam | PF02885 | Glycosyl transferase family, helical bundle domain | IPR017459 | Glycosyl transferase family 3, N-terminal domain | 6 | 67 | 4.7E-11 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.