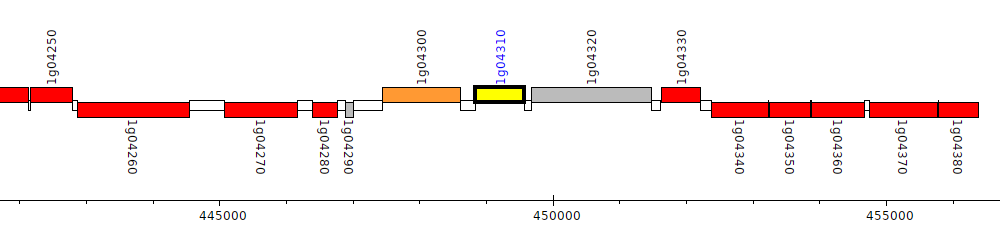
Burkholderia glumae BGR1, bglu_1g04310
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0045454 | cell redox homeostasis |
Inferred from Sequence Model
Term mapped from: InterPro:PS00194
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0042597 | periplasmic space |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.10.450.70
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF54423 | IPR009094 | Disulphide bond isomerase DsbC/G, N-terminal domain superfamily | 35 | 88 | 5.49E-12 | |
| Gene3D | G3DSA:3.10.450.70 | IPR009094 | Disulphide bond isomerase DsbC/G, N-terminal domain superfamily | 24 | 89 | 1.1E-9 | |
| Pfam | PF13098 | Thioredoxin-like domain | IPR012336 | Thioredoxin-like fold | 114 | 237 | 5.9E-32 |
| Gene3D | G3DSA:3.40.30.10 | 106 | 240 | 9.8E-39 | |||
| ProSitePatterns | PS00194 | Thioredoxin family active site. | IPR017937 | Thioredoxin, conserved site | 122 | 140 | - |
| CDD | cd03020 | DsbA_DsbC_DsbG | IPR033954 | Disulphide bond isomerase, DsbC/G | 43 | 237 | 3.17536E-88 |
| ProSiteProfiles | PS51257 | Prokaryotic membrane lipoprotein lipid attachment site profile. | 1 | 21 | 6.0 | ||
| SUPERFAMILY | SSF52833 | IPR036249 | Thioredoxin-like superfamily | 84 | 237 | 1.33E-37 | |
| Pfam | PF10411 | Disulfide bond isomerase protein N-terminus | IPR018950 | Disulphide bond isomerase, DsbC/G, N-terminal | 33 | 86 | 7.6E-19 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.